R packages: Difference between revisions
| Line 1,598: | Line 1,598: | ||
* [http://www.markvanderloo.eu/yaRb/2019/01/11/add-a-static-pdf-vignette-to-an-r-package/ Add a static pdf vignette to an R package] | * [http://www.markvanderloo.eu/yaRb/2019/01/11/add-a-static-pdf-vignette-to-an-r-package/ Add a static pdf vignette to an R package] | ||
* [https://www.rdocumentation.org/packages/devtools/versions/2.2.0/topics/build_vignettes devtools::build_vignettes()]. The files are copied in the 'doc' directory and an vignette index is created in 'Meta/vignette.rds', as they would be in a built package. | * [https://www.rdocumentation.org/packages/devtools/versions/2.2.0/topics/build_vignettes devtools::build_vignettes()]. The files are copied in the 'doc' directory and an vignette index is created in 'Meta/vignette.rds', as they would be in a built package. | ||
* [https://stackoverflow.com/ | * [https://stackoverflow.com/a/37692810 build_vignette can't find functions] | ||
== README.Rmd & README.md files == | == README.Rmd & README.md files == | ||
Revision as of 08:08, 30 September 2019
R package management
Packages loaded at startup
getOption("defaultPackages")
How to install a new package
utils::install.packages()
- By default, install.packages() will check versions and install uninstalled packages shown in 'Depends', 'Imports' , and 'LinkingTo' fields. See R-exts manual.
- If we want to install packages listed in 'Suggests' field, we should specify it explicitly by using dependencies argument:
install.packages(XXXX, dependencies = c("Depends", "Imports", "Suggests", "LinkingTo")) # OR install.packages(XXXX, dependencies = TRUE)For example, if I use a plain install.packages() command to install downloader package, it only installs 'digest' and 'downloader' packages. If I useinstall.packages("downloader", dependencies=TRUE)it will also install 'testhat' package.
- Even a warning is given when some imports package is not available, it does not stop the installation (trying to install plotROC package on mac R 3.5.3). See also Warning: dependency ‘XXX’ is not available.
> install.packages("plotROC") --- Please select a CRAN mirror for use in this session --- Warning: dependency ‘gridSVG’ is not available trying URL 'https://cloud.r-project.org/bin/macosx/el-capitan/contrib/3.5/plotROC_2.2.1.tgz' Content type 'application/x-gzip' length 1288370 bytes (1.2 MB) ================================================== downloaded 1.2 MB The downloaded binary packages are in /var/folders/2q/slryb0rx4tj97t66v7l6pwvr_z6g3s/T//Rtmptrf2ZS/downloaded_packages > library(gridSVG) Error in library(gridSVG) : there is no package called ‘gridSVG’ > install.packages("gridSVG", type = "source") Warning message: package ‘gridSVG’ is not available (for R version 3.5.3) > ap <- available.packages() > "gridSVG" %in% rownames(ap) # [1] FALSE > dim(ap) # [1] 14220 17 > library(plotROC) Loading required package: ggplot2
- Look at CRAN, we see the latest version (1.7-0) of gridSVG depends on the latest version of R (3.6.0). But why even the source is removed from the old version of R?
- The install.packages function source code can be found in R -> src -> library -> utils -> R -> packages2.R file from Github repository (put 'install.packages' in the search box).
pacman: Install and load the package at the same time, installing temporarily
p_load() function from the pacman package.
An example of installing the purrr package.
Also used by BBC Visual and Data Journalism cookbook for R graphics
Also used by Biowulf/NIH
remotes
Download and install R packages stored in 'GitHub', 'BitBucket', or plain 'subversion' or 'git' repositories. This package is a lightweight replacement of the 'install_*' functions in 'devtools'.
- https://cran.r-project.org/web/packages/remotes/index.html
- https://www.rdocumentation.org/packages/remotes/versions/2.0.2
To install a package from a local machines with all dependency, run remotes::install_local(path = "MyPackage.tar.gz", dependencies=TRUE) or devtools::install() though the later requires to untar the source first.
devtools
pak
https://cran.r-project.org/web/packages/pak/index.html
BiocManager
https://cran.r-project.org/web/packages/BiocManager/index.html
pkgman
pkgman: A fresh approach to package installation - Gábor Csárdi
Check installed Bioconductor version
Following this post, use tools:::.BioC_version_associated_with_R_version().
Mind the '.' in front of the 'BioC'. It may be possible for some installed packages to have been sourced from a different BioC version.
tools:::.BioC_version_associated_with_R_version() # `3.6' tools:::.BioC_version_associated_with_R_version() == '3.6' # TRUE
CRAN Package Depends on Bioconductor Package
For example, if I run install.packages("NanoStringNorm") to install the package from CRAN, I may get
ERROR: dependency ‘vsn’ is not available for package ‘NanoStringNorm’
This is because the NanoStringNorm package depends on the vsn package which is on Bioconductor.
Another instance is CRAN's 'biospear' (actually 'plsRcox') depends on Bioc's 'survcomp' & 'mixOmics'.
One solution is to run a line setRepositories(ind=1:2). See this post or this one. Note that the default repository list can be found at (Ubuntu) /usr/lib/R/etc/repositories file.
options("repos") # display the available repositories (only CRAN)
setRepositories(ind=1:2)
options("repos") # CRAN and bioc are included
# CRAN
# "https://cloud.r-project.org"
# "https://bioconductor.org/packages/3.6/bioc"
install.packages("biospear") # it will prompt to select CRAN
install.packages("biospear", repos = "http://cran.rstudio.com") # NOT work since bioc repos is erased
This will also install the BiocInstaller package if it has not been installed before. See also Install Bioconductor Packages.
install a tar.gz (e.g. an archived package) from a local directory
R CMD INSTALL <package-name>.tar.gz
Or in R:
# Method 1: cannot install dependencies
install.packages(<pathtopackage>, repos = NULL)
# These paths can be source directories or archives or binary package
# archive files (as created by ‘R CMD build --binary’).
# (‘http://’ and ‘file://’ URLs are also accepted and the files
# will be downloaded and installed from local copies.)
# Method 2: take care of dependencies from CRAN
devtools::install(<directory to package>, dependencies = TRUE)
# this will use 'R CMD INSTALL' to install the package.
# It will try to install dependencies of the package from CRAN,
# if they're not already installed.
The installation process can be nasty due to the dependency issue. Consider the 'biospear' package
biospear - plsRcox (archived) - plsRglm (archived) - bipartite
- lars
- pls
- kernlab
- mixOmics (CRAN->Bioconductor)
- risksetROC
- survcomp (Bioconductor)
- rms
So in order to install the 'plsRcox' package, we need to do the following steps. Note: plsRcox package is back on 6/2/2018.
# For curl
system("apt update")
system("apt install curl libcurl4-openssl-dev libssl-dev")
# For X11
system("apt install libcgal-dev libglu1-mesa-dev libglu1-mesa-dev")
system("apt install libfreetype6-dev") # https://stackoverflow.com/questions/31820865/error-in-installing-rgl-package
source("https://bioconductor.org/biocLite.R")
biocLite("survcomp") # this has to be run before the next command of installing a bunch of packages from CRAN
install.packages("https://cran.r-project.org/src/contrib/Archive/biospear/biospear_1.0.1.tar.gz",
repos = NULL, type="source")
# ERROR: dependencies ‘pkgconfig’, ‘cobs’, ‘corpcor’, ‘devtools’, ‘glmnet’, ‘grplasso’, ‘mboost’, ‘plsRcox’,
# ‘pROC’, ‘PRROC’, ‘RCurl’, ‘survAUC’ are not available for package ‘biospear’
install.packages(c("pkgconfig", "cobs", "corpcor", "devtools", "glmnet", "grplasso", "mboost",
"plsRcox", "pROC", "PRROC", "RCurl", "survAUC"))
# optional: install.packages(c("doRNG", "mvnfast"))
install.packages("https://cran.r-project.org/src/contrib/Archive/biospear/biospear_1.0.1.tar.gz",
repos = NULL, type="source")
# OR
# devtools::install_github("cran/biospear")
library(biospear) # verify
To install the (deprecated, bioc) packages 'inSilicoMerging',
biocLite(c('rjson', 'Biobase', 'RCurl'))
# destination directory is required
# download.file("http://www.bioconductor.org/packages/3.3/bioc/src/contrib/inSilicoDb_2.7.0.tar.gz",
# "~/Downloads/inSilicoDb_2.7.0.tar.gz")
# download.file("http://www.bioconductor.org/packages/3.3/bioc/src/contrib/inSilicoMerging_1.15.0.tar.gz",
# "~/Downloads/inSilicoMerging_1.15.0.tar.gz")
# ~/Downloads or $HOME/Downloads won't work in untar()
# untar("~/Downloads/inSilicoDb_2.7.0.tar.gz", exdir="/home/brb/Downloads")
# untar("~/Downloads/inSilicoMerging_1.15.0.tar.gz", exdir="/home/brb/Downloads")
# install.packages("~/Downloads/inSilicoDb", repos = NULL)
# install.packages("~/Downloads/inSilicoMerging", repos = NULL)
install.packages("http://www.bioconductor.org/packages/3.3/bioc/src/contrib/inSilicoDb_2.7.0.tar.gz",
repos = NULL, type = "source")
install.packages("http://www.bioconductor.org/packages/3.3/bioc/src/contrib/inSilicoMerging_1.15.0.tar.gz",
repos = NULL, type = "source")
Install multiple versions of the same R package
https://stackoverflow.com/a/2989369
library(foo, lib.loc="~/dev/foo/v1") ## loads v1 # and library(foo, lib.loc="~/dev/foo/v2") ## loads v2
The same works for install.packages(). help(install.packages)
The install_version() from devtools and remotes will overwrite the existing installation.
Query an R package installed locally
packageDescription("MASS")
packageVersion("MASS")
Query an R package (from CRAN) basic information
packageStatus() # Summarize information about installed packages available.packages() # List Available Packages at CRAN-like Repositories
The available.packages() command is useful for understanding package dependency. Use setRepositories() or 'RGUI -> Packages -> select repositories' to select repositories and options()$repos to check or change the repositories.
Also the packageStatus() is another useful function for query how many packages are in the repositories, how many have been installed, and individual package status (installed or not, needs to be upgraded or not).
> options()$repos
CRAN
"https://cran.rstudio.com/"
> packageStatus()
Number of installed packages:
ok upgrade unavailable
C:/Program Files/R/R-3.0.1/library 110 0 1
Number of available packages (each package counted only once):
installed not installed
http://watson.nci.nih.gov/cran_mirror/bin/windows/contrib/3.0 76 4563
http://www.stats.ox.ac.uk/pub/RWin/bin/windows/contrib/3.0 0 5
http://www.bioconductor.org/packages/2.12/bioc/bin/windows/contrib/3.0 16 625
http://www.bioconductor.org/packages/2.12/data/annotation/bin/windows/contrib/3.0 4 686
> tmp <- available.packages()
> str(tmp)
chr [1:5975, 1:17] "A3" "ABCExtremes" "ABCp2" "ACCLMA" "ACD" "ACNE" "ADGofTest" "ADM3" "AER" ...
- attr(*, "dimnames")=List of 2
..$ : chr [1:5975] "A3" "ABCExtremes" "ABCp2" "ACCLMA" ...
..$ : chr [1:17] "Package" "Version" "Priority" "Depends" ...
> tmp[1:3,]
Package Version Priority Depends Imports LinkingTo Suggests
A3 "A3" "0.9.2" NA "xtable, pbapply" NA NA "randomForest, e1071"
ABCExtremes "ABCExtremes" "1.0" NA "SpatialExtremes, combinat" NA NA NA
ABCp2 "ABCp2" "1.1" NA "MASS" NA NA NA
Enhances License License_is_FOSS License_restricts_use OS_type Archs MD5sum NeedsCompilation File
A3 NA "GPL (>= 2)" NA NA NA NA NA NA NA
ABCExtremes NA "GPL-2" NA NA NA NA NA NA NA
ABCp2 NA "GPL-2" NA NA NA NA NA NA NA
Repository
A3 "http://watson.nci.nih.gov/cran_mirror/bin/windows/contrib/3.0"
ABCExtremes "http://watson.nci.nih.gov/cran_mirror/bin/windows/contrib/3.0"
ABCp2 "http://watson.nci.nih.gov/cran_mirror/bin/windows/contrib/3.0"
And the following commands find which package depends on Rcpp and also which are from bioconductor repository.
> pkgName <- "Rcpp"
> rownames(tmp)[grep(pkgName, tmp[,"Depends"])]
> tmp[grep("Rcpp", tmp[,"Depends"]), "Depends"]
> ind <- intersect(grep(pkgName, tmp[,"Depends"]), grep("bioconductor", tmp[, "Repository"]))
> rownames(grep)[ind]
NULL
> rownames(tmp)[ind]
[1] "ddgraph" "DESeq2" "GeneNetworkBuilder" "GOSemSim" "GRENITS"
[6] "mosaics" "mzR" "pcaMethods" "Rdisop" "Risa"
[11] "rTANDEM"
CRAN vs Bioconductor packages
> R.version # 3.4.3
# CRAN
> x <- available.packages()
> dim(x)
[1] 12581 17
# Bioconductor Soft
> biocinstallRepos()
BioCsoft
"https://bioconductor.org/packages/3.6/bioc"
BioCann
"https://bioconductor.org/packages/3.6/data/annotation"
BioCexp
"https://bioconductor.org/packages/3.6/data/experiment"
CRAN
"https://cran.rstudio.com/"
> y <- available.packages(repos = biocinstallRepos()[1])
> dim(y)
[1] 1477 17
> intersect(x[, "Package"], y[, "Package"])
character(0)
# Bioconductor Annotation
> dim(available.packages(repos = biocinstallRepos()[2]))
[1] 909 17
# Bioconductor Experiment
> dim(available.packages(repos = biocinstallRepos()[3]))
[1] 326 17
# CRAN + All Bioconductor
> z <- available.packages(repos = biocinstallRepos())
> dim(z)
[1] 15292 17
Downloading Bioconductor package with an old R
When I try to download the GenomicDataCommons package using R 3.4.4 with Bioc 3.6 (the current R version is 3.5.0), it was found it can only install version 1.2.0 instead the latest version 1.4.1.
It does not work by running biocLite("BiocUpgrade") to upgrade Bioc from 3.6 to 3.7.
source("https://bioconductor.org/biocLite.R")
biocLite("BiocUpgrade")
# Error: Bioconductor version 3.6 cannot be upgraded with R version 3.4.4
Analyzing data on CRAN packages
New undocumented function in R 3.4.0: tools::CRAN_package_db()
http://blog.revolutionanalytics.com/2017/05/analyzing-data-on-cran-packages.html
R package location when they are installed by root
/usr/local/lib/R/site-library
R_LIBS_USER is empty in R 3.4.1
See install.package() error, R_LIBS_USER is empty in R 3.4.1.
Add a personal directory to .libPaths()
.libPaths( c( .libPaths(), "~/userLibrary") )
No need to use the assignment operator.
Install personal R packages after upgrade R, .libPaths(), Rprofile.site, R_LIBS_USER
Scenario: We already have installed many R packages under R 3.1.X in the user's directory. Now we upgrade R to a new version (3.2.X). We like to have these packages available in R 3.2.X.
For Windows OS, refer to R for Windows FAQ
The follow method works on Linux and Windows.
Make sure only one instance of R is running
# Step 1. update R's built-in packages and install them on my personal directory
update.packages(ask=FALSE, checkBuilt = TRUE, repos="http://cran.rstudio.com")
# Step 2. update Bioconductor packages
.libPaths() # The first one is my personal directory
# [1] "/home/brb/R/x86_64-pc-linux-gnu-library/3.2"
# [2] "/usr/local/lib/R/site-library"
# [3] "/usr/lib/R/site-library"
# [4] "/usr/lib/R/library"
Sys.getenv("R_LIBS_USER") # may or may not equivalent to .libPaths()[1]
ul <- unlist(strsplit(Sys.getenv("R_LIBS_USER"), "/"))
src <- file.path(paste(ul[1:(length(ul)-1)], collapse="/"), "3.1")
des <- file.path(paste(ul[1:(length(ul)-1)], collapse="/"), "3.2")
pkg <- dir(src, full.names = TRUE)
if (!file.exists(des)) dir.create(des) # If 3.2 subdirectory does not exist yet!
file.copy(pkg, des, overwrite=FALSE, recursive = TRUE)
source("http://www.bioconductor.org/biocLite.R")
biocLite(ask = FALSE)
From Robert Kabacoff (R in Action)
- If you have a customized Rprofile.site file (see appendix B), save a copy outside of R.
- Launch your current version of R and issue the following statements
oldip <- installed.packages()[,1] save(oldip, file="path/installedPackages.Rdata")
where path is a directory outside of R.
- Download and install the newer version of R.
- If you saved a customized version of the Rprofile.site file in step 1, copy it into the new installation.
- Launch the new version of R, and issue the following statements
load("path/installedPackages.Rdata")
newip <- installed.packages()[,1]
for(i in setdiff(oldip, newip))
install.packages(i)
where path is the location specified in step 2.
- Delete the old installation (optional).
This approach will install only packages that are available from the CRAN. It won’t find packages obtained from other locations. In fact, the process will display a list of packages that can’t be installed For example for packages obtained from Bioconductor, use the following method to update packages
source(http://bioconductor.org/biocLite.R) biocLite(PKGNAME)
Would you like to use a personal library instead?
Some posts from internet
- Setting R_LIBS & avoiding “Would you like to use a personal library instead?”. Note: I try to create ~/.Renviron to add my personal folder in it. But update.packages() still asks me if I like to use a personal library instead (tested on Ubuntu + R 3.4).
- automatically create personal library in R. Using suppressUpdates + specify lib in biocLite() or update.packages(Sys.getenv("R_LIBS_USER"), ask = F)
# create local user library path (not present by default) dir.create(path = Sys.getenv("R_LIBS_USER"), showWarnings = FALSE, recursive = TRUE) # install to local user library path install.packages(p, lib = Sys.getenv("R_LIBS_USER"), repos = "https://cran.rstudio.com/") # Bioconductor version biocLite(p, suppressUpdates = TRUE, lib = Sys.getenv("R_LIBS_USER"))
The problem can happen if the R was installed to the C:\Program Files\R folder by users but then some main packages want to be upgraded. R will always pops a message 'Would you like to use a personal library instead?'.
To suppress the message and use the personal library always,
- Run R as administrator. If you do that, main packages can be upgraded from C:\Program Files\R\R-X.Y.Z\library folder.
- Writable R package directory cannot be found and a this. A solution here is to change the security of the R library folder so the user has a full control on the folder.
- Does R run under Windows Vista/7/8/Server 2008? There are 3 ways to get around the issue.
- I don’t have permission to write to the R-3.3.2\library directory
Actually the following hints will help us to create a convenient function UpdateMainLibrary() which will install updated main packages in the user's Documents directory without a warning dialog.
- .libPaths() only returns 1 string "C:/Program Files/R/R-x.y.z/library" on the machines that does not have this problem
- .libPaths() returns two strings "C:/Users/USERNAME/Documents/R/win-library/x.y" & "C:/Program Files/R/R-x.y.z/library" on machines with the problem.
UpdateMainLibrary <- function() {
# Update main/site packages
# The function is used to fix the problem 'Would you like to use a personal library instead?'
if (length(.libPaths()) == 1) return()
ind_mloc <- grep("Program", .libPaths()) # main library e.g. 2
ind_ploc <- grep("Documents", .libPaths()) # personal library e.g. 1
if (length(ind_mloc) > 0L && length(ind_ploc) > 0L)
# search outdated main packages
old_mloc <- ! old.packages(.libPaths()[ind_mloc])[, "Package"] %in%
installed.packages(.libPaths()[ind_ploc])[, "Package"]
oldpac <- old.packages(.libPaths()[ind_mloc])[old_mloc, "Package"]
if (length(oldpac) > 0L)
install.packages(oldpac, .libPaths()[ind_ploc])
}
On Linux,
> update.packages()
...
The downloaded source packages are in
‘/tmp/RtmpBrYccd/downloaded_packages’
Warning in install.packages(update[instlib == l, "Package"], l, contriburl = contriburl, :
'lib = "/opt/R/3.5.0/lib/R/library"' is not writable
Would you like to use a personal library instead? (yes/No/cancel) yes
...
> system("ls -lt /home/brb/R/x86_64-pc-linux-gnu-library/3.5 | head")
total 224
drwxrwxr-x 9 brb brb 4096 Oct 3 09:30 survival
drwxrwxr-x 9 brb brb 4096 Oct 3 09:29 mgcv
drwxrwxr-x 10 brb brb 4096 Oct 3 09:29 MASS
drwxrwxr-x 9 brb brb 4096 Oct 3 09:29 foreign
# So new versions of survival, mgc, MASS, foreign are installed in the personal directory
# The update.packages() will issue warnings if we try to run it again.
# It's OK to ignore these warnings.
> update.packages()
Warning: package 'foreign' in library '/opt/R/3.5.0/lib/R/library' will not be updated
Warning: package 'MASS' in library '/opt/R/3.5.0/lib/R/library' will not be updated
Warning: package 'mgcv' in library '/opt/R/3.5.0/lib/R/library' will not be updated
Warning: package 'survival' in library '/opt/R/3.5.0/lib/R/library' will not be updated
List vignettes from a package
vignette(package=PACKAGENAME)
List data from a package
data(package=PACKAGENAME)
List installed packages and versions
- http://heuristicandrew.blogspot.com/2015/06/list-of-user-installed-r-packages-and.html
- checkpoint package
ip <- as.data.frame(installed.packages()[,c(1,3:4)]) rownames(ip) <- NULL unique(ip$Priority) # [1] <NA> base recommended # Levels: base recommended ip <- ip[is.na(ip$Priority),1:2,drop=FALSE] print(ip, row.names=FALSE)
Query the names of outdated packages
psi <- packageStatus()$inst subset(psi, Status == "upgrade", drop = FALSE) # Package LibPath Version Priority Depends # RcppArmadillo RcppArmadillo C:/Users/brb/Documents/R/win-library/3.2 0.5.100.1.0 <NA> <NA> # Matrix Matrix C:/Program Files/R/R-3.2.0/library 1.2-0 recommended R (>= 2.15.2), methods # Imports LinkingTo Suggests # RcppArmadillo Rcpp (>= 0.11.0) Rcpp RUnit, Matrix, pkgKitten # Matrix graphics, grid, stats, utils, lattice <NA> expm, MASS # Enhances License License_is_FOSS License_restricts_use OS_type MD5sum # RcppArmadillo <NA> GPL (>= 2) <NA> <NA> <NA> <NA> # Matrix MatrixModels, graph, SparseM, sfsmisc GPL (>= 2) <NA> <NA> <NA> <NA> # NeedsCompilation Built Status # RcppArmadillo yes 3.2.0 upgrade # Matrix yes 3.2.0 upgrade
The above output does not show the package version from the latest packages on CRAN. So the following snippet does that.
psi <- packageStatus()$inst
pl <- unname(psi$Package[psi$Status == "upgrade"]) # List package names
ap <- as.data.frame(available.packages()[, c(1,2,3)], stringsAsFactors = FALSE)
out <- cbind(subset(psi, Status == "upgrade")[, c("Package", "Version")], ap[match(pl, ap$Package), "Version"])
colnames(out)[2:3] <- c("OldVersion", "NewVersion")
rownames(out) <- NULL
out
# Package OldVersion NewVersion
# 1 RcppArmadillo 0.5.100.1.0 0.5.200.1.0
# 2 Matrix 1.2-0 1.2-1
To consider also the packages from Bioconductor, we have the following code. Note that "3.1" means the Bioconductor version and "3.2" is the R version. See Bioconductor release versions page.
psic <- packageStatus(repos = c(contrib.url(getOption("repos")),
"http://bioconductor.org/packages/3.1/bioc/bin/windows/contrib/3.2",
"http://www.bioconductor.org/packages/3.1/data/annotation/bin/windows/contrib/3.2"))$inst
subset(psic, Status == "upgrade", drop = FALSE)
pl <- unname(psic$Package[psic$Status == "upgrade"])
ap <- as.data.frame(available.packages(c(contrib.url(getOption("repos")),
"http://bioconductor.org/packages/3.1/bioc/bin/windows/contrib/3.2",
"http://www.bioconductor.org/packages/3.1/data/annotation/bin/windows/contrib/3.2"))[, c(1:3)],
stringAsFactors = FALSE)
out <- cbind(subset(psic, Status == "upgrade")[, c("Package", "Version")], ap[match(pl, ap$Package), "Version"])
colnames(out)[2:3] <- c("OldVersion", "NewVersion")
rownames(out) <- NULL
out
# Package OldVersion NewVersion
# 1 limma 3.24.5 3.24.9
# 2 RcppArmadillo 0.5.100.1.0 0.5.200.1.0
# 3 Matrix 1.2-0 1.2-1
Searching for packages in CRAN
- pkgsearch package - Search R packages on CRAN
- Fishing for packages in CRAN
- CRAN now has 10,000 R packages. Here's how to find the ones you need
- Searching for R packages, packagefinder package
library(packagefinder) findPackage("survival") # 272 out of 13256 CRAN packages found in 5 seconds findPackage("follic") # No results found. # Actually 'follic' comes from randomForestSRC package # https://www.rdocumentation.org/packages/randomForestSRC/versions/2.7.0The result is shown in an html format with columns of SCORE, NAME, DESC_SHORT, DOWNL_TOTAL & GO.
METACRAN (www.r-pkg.org) - Search and browse all CRAN/R packages
- Source code on https://github.com/metacran. The 'PACKAGES' file is updated regularly to Github.
- Announcement on R/mailing list
- Author's homepage on http://gaborcsardi.org/.
cranly visualisations and summaries for R packages
Exploring R packages with cranly
Query top downloaded packages, download statistics
- Daily download statistics http://cran-logs.rstudio.com/. Note the page is split into 'package' download and 'R' download. It tracks
- Package: date, time, size, r_version, r_arch, r_os, package, version, country, ip_id.
- R: date, time, size, R version, os (win/src/osx), county, ip_id (reset daily).
- Original methods
- https://www.r-bloggers.com/finally-tracking-cran-packages-downloads/. The code still works.
- My modified code for showing the top download R packages. The original code suffers from memory issue (tested on my 64GB Linux box) when it is calling rbindlist() from data.table package with large data. Still 64GB is required since the matrix is 369M by 4 (12GB).
- Popularity bigdata / large data packages in R and ffbase useR presentation
- cranlogs package - Download Logs from the RStudio CRAN Mirror
- 2018 through {cranlogs}
- Shiny app by Hadley (works for packages on CRAN only)
- cranlogs 2.1.1 is on CRAN! 5/2/2019
- packageRank package: Computing and Visualizing CRAN Downloads
- pkginfo: Tools for Retrieving R Package Information
- For Bioconductor packages, try BiocPkgTools. See the paper.
installation path not writeable from running biocLite()
When I ran biocLite() to install a new package, I got a message (the Bioc packages are installed successfully anyway)
... * DONE (curatedOvarianData) The downloaded source packages are in ‘/tmp/RtmpHxnH2K/downloaded_packages’ installation path not writeable, unable to update packages: rgl, rJava, codetools, foreign, lattice, MASS, spatial, survival
However, if I uses install.package() it can update the package
> packageVersion("survival")
[1] ‘2.42.3’
> update.packages("survival") # Not working though no error message
> packageVersion("survival")
[1] ‘2.42.3’
> install.packages("survival")
Installing package into ‘/home/brb/R/x86_64-pc-linux-gnu-library/3.4’
...
* DONE (survival)
The downloaded source packages are in
‘/tmp/RtmpHxnH2K/downloaded_packages’
> packageVersion("survival")
[1] ‘2.42.6’
> library(survival)
> sessionInfo() # show survival package 2.42-6 was attached
It makes sense to always use personal directory when we install packages. See .libPaths().
Warning: cannot remove prior installation of package
Instance 1.
# Install the latest hgu133plus2cdf package
# Remove/Uninstall hgu133plus2.db package
# Put/Install an old version of IRanges (eg version 1.18.2 while currently it is version 1.18.3)
# Test on R 3.0.1
library(hgu133plus2cdf) # hgu133pluscdf does not depend or import IRanges
source("http://bioconductor.org/biocLite.R")
biocLite("hgu133plus2.db", ask=FALSE) # hgu133plus2.db imports IRanges
# Warning:cannot remove prior installation of package 'IRanges'
# Open Windows Explorer and check IRanges folder. Only see libs subfolder.
Note:
- In the above example, all packages were installed under C:\Program Files\R\R-3.0.1\library\.
- In another instance where I cannot reproduce the problem, new R packages were installed under C:\Users\xxx\Documents\R\win-library\3.0\. The different thing is IRanges package CAN be updated but if I use packageVersion("IRanges") command in R, it still shows the old version.
- The above were tested on a desktop.
Instance 2.
# On a fresh R 3.2.0, I install Bioconductor's depPkgTools & lumi packages. Then I close R, re-open it,
# and install depPkgTools package again.
> source("http://bioconductor.org/biocLite.R")
Bioconductor version 3.1 (BiocInstaller 1.18.2), ?biocLite for help
> biocLite("pkgDepTools")
BioC_mirror: http://bioconductor.org
Using Bioconductor version 3.1 (BiocInstaller 1.18.2), R version 3.2.0.
Installing package(s) ‘pkgDepTools’
trying URL 'http://bioconductor.org/packages/3.1/bioc/bin/windows/contrib/3.2/pkgDepTools_1.34.0.zip'
Content type 'application/zip' length 390579 bytes (381 KB)
downloaded 381 KB
package ‘pkgDepTools’ successfully unpacked and MD5 sums checked
Warning: cannot remove prior installation of package ‘pkgDepTools’
The downloaded binary packages are in
C:\Users\brb\AppData\Local\Temp\RtmpYd2l7i\downloaded_packages
> library(pkgDepTools)
Error in library(pkgDepTools) : there is no package called ‘pkgDepTools’
The pkgDepTools library folder appears in C:\Users\brb\Documents\R\win-library\3.2, but it is empty. The weird thing is if I try the above steps again, I cannot reproduce the problem.
Warning: dependency ‘XXX’ is not available
How should I deal with “package 'xxx' is not available (for R version x.y.z)” warning?
Error: there is no package called XXX
The error happened when I try to run library() command on a package that was just installed. R 3.6.0. 'biospear' version is 1.0.2. macOS.
> library(biospear) Loading required package: pkgconfig Error: package or namespace load failed for ‘biospear’ in loadNamespace(i, c(lib.loc, .libPaths()), versionCheck = vI[[i]]): there is no package called ‘mixOmics’
Note
- The package mixOmics was removed from CRAN. It is now available on Bioconductor.
- Tested to install on a docker container: docker run --net=host -it --rm r-base
ERROR: dependency ‘car’ is not available for package ‘plsRglm’ * removing ‘/usr/local/lib/R/site-library/plsRglm’ ERROR: dependencies ‘plsRglm’, ‘mixOmics’, ‘survcomp’ are not available for package ‘plsRcox’ * removing ‘/usr/local/lib/R/site-library/plsRcox’ ERROR: dependencies ‘devtools’, ‘plsRcox’, ‘RCurl’ are not available for package ‘biospear’ * removing ‘/usr/local/lib/R/site-library/biospear’
The car package looks OK on CRAN. survcomp was moved from CRAN to Bioconductor too. - As we can see above, the official r-base image does not contain libraries enough to install RCurl/devtools packages. Consider the tidyverse image (based on RStudio image) from the rocker project.
docker pull rocker/tidyverse:3.6.0 docker run --net=host -it --rm -e PASSWORD=password -p 8787:8787 rocker/tidyverse:3.6.0 # the default username is 'rstudio' # Open a browser, log in. Run 'install.packages("RCurl")'. It works. - Testing on Mint linux also shows errors about dependencies of mixOmics and survcomp.
- The best practice to install a package that may depend on packages located/moved to Bioconductor: Run setRepositories(ind=1:2) before calling install.packages(). However, it does not remedy the situation that the 1st level imports package (eg plsRcox) was installed before but the 2nd level imports package (eg mixOmics) was not installed.
- dependsOnPkgs(): Find Reverse Dependencies
Warning: Unable to move temporary installation
The problem seems to happen only on virtual machines (Virtualbox).
- Warning: unable to move temporary installation `C:\Users\brb\Documents\R\win-library\3.0\fileed8270978f5\quadprog` to `C:\Users\brb\Documents\R\win-library\3.0\quadprog` when I try to run 'install.packages("forecast").
- Warning: unable to move temporary installation ‘C:\Users\brb\Documents\R\win-library\3.2\file5e0104b5b49\plyr’ to ‘C:\Users\brb\Documents\R\win-library\3.2\plyr’ when I try to run 'biocLite("lumi")'. The other dependency packages look fine although I am not sure if any unknown problem can happen (it does, see below).
Here is a note of my trouble shooting.
- If I try to ignore the warning and load the lumi package. I will get an error.
- If I try to run biocLite("lumi") again, it will only download & install lumi without checking missing 'plyr' package. Therefore, when I try to load the lumi package, it will give me an error again.
- Even I install the plyr package manually, library(lumi) gives another error - missing mclust package.
> biocLite("lumi")
trying URL 'http://bioconductor.org/packages/3.1/bioc/bin/windows/contrib/3.2/BiocInstaller_1.18.2.zip'
Content type 'application/zip' length 114097 bytes (111 KB)
downloaded 111 KB
...
package ‘lumi’ successfully unpacked and MD5 sums checked
The downloaded binary packages are in
C:\Users\brb\AppData\Local\Temp\RtmpyUjsJD\downloaded_packages
Old packages: 'BiocParallel', 'Biostrings', 'caret', 'DESeq2', 'gdata', 'GenomicFeatures', 'gplots', 'Hmisc', 'Rcpp', 'RcppArmadillo', 'rgl',
'stringr'
Update all/some/none? [a/s/n]: a
also installing the dependencies ‘Rsamtools’, ‘GenomicAlignments’, ‘plyr’, ‘rtracklayer’, ‘gridExtra’, ‘stringi’, ‘magrittr’
trying URL 'http://bioconductor.org/packages/3.1/bioc/bin/windows/contrib/3.2/Rsamtools_1.20.1.zip'
Content type 'application/zip' length 8138197 bytes (7.8 MB)
downloaded 7.8 MB
...
package ‘Rsamtools’ successfully unpacked and MD5 sums checked
package ‘GenomicAlignments’ successfully unpacked and MD5 sums checked
package ‘plyr’ successfully unpacked and MD5 sums checked
Warning: unable to move temporary installation ‘C:\Users\brb\Documents\R\win-library\3.2\file5e0104b5b49\plyr’
to ‘C:\Users\brb\Documents\R\win-library\3.2\plyr’
package ‘rtracklayer’ successfully unpacked and MD5 sums checked
package ‘gridExtra’ successfully unpacked and MD5 sums checked
package ‘stringi’ successfully unpacked and MD5 sums checked
package ‘magrittr’ successfully unpacked and MD5 sums checked
package ‘BiocParallel’ successfully unpacked and MD5 sums checked
package ‘Biostrings’ successfully unpacked and MD5 sums checked
Warning: cannot remove prior installation of package ‘Biostrings’
package ‘caret’ successfully unpacked and MD5 sums checked
package ‘DESeq2’ successfully unpacked and MD5 sums checked
package ‘gdata’ successfully unpacked and MD5 sums checked
package ‘GenomicFeatures’ successfully unpacked and MD5 sums checked
package ‘gplots’ successfully unpacked and MD5 sums checked
package ‘Hmisc’ successfully unpacked and MD5 sums checked
package ‘Rcpp’ successfully unpacked and MD5 sums checked
package ‘RcppArmadillo’ successfully unpacked and MD5 sums checked
package ‘rgl’ successfully unpacked and MD5 sums checked
package ‘stringr’ successfully unpacked and MD5 sums checked
The downloaded binary packages are in
C:\Users\brb\AppData\Local\Temp\RtmpyUjsJD\downloaded_packages
> library(lumi)
Error in loadNamespace(i, c(lib.loc, .libPaths()), versionCheck = vI[[i]]) :
there is no package called ‘plyr’
Error: package or namespace load failed for ‘lumi’
> search()
[1] ".GlobalEnv" "package:BiocInstaller" "package:Biobase" "package:BiocGenerics" "package:parallel" "package:stats"
[7] "package:graphics" "package:grDevices" "package:utils" "package:datasets" "package:methods" "Autoloads"
[13] "package:base"
> biocLite("lumi")
BioC_mirror: http://bioconductor.org
Using Bioconductor version 3.1 (BiocInstaller 1.18.2), R version 3.2.0.
Installing package(s) ‘lumi’
trying URL 'http://bioconductor.org/packages/3.1/bioc/bin/windows/contrib/3.2/lumi_2.20.1.zip'
Content type 'application/zip' length 18185326 bytes (17.3 MB)
downloaded 17.3 MB
package ‘lumi’ successfully unpacked and MD5 sums checked
The downloaded binary packages are in
C:\Users\brb\AppData\Local\Temp\RtmpyUjsJD\downloaded_packages
> search()
[1] ".GlobalEnv" "package:BiocInstaller" "package:Biobase" "package:BiocGenerics" "package:parallel" "package:stats"
[7] "package:graphics" "package:grDevices" "package:utils" "package:datasets" "package:methods" "Autoloads"
[13] "package:base"
> library(lumi)
Error in loadNamespace(i, c(lib.loc, .libPaths()), versionCheck = vI[[i]]) :
there is no package called ‘plyr’
Error: package or namespace load failed for ‘lumi’
> biocLite("plyr")
BioC_mirror: http://bioconductor.org
Using Bioconductor version 3.1 (BiocInstaller 1.18.2), R version 3.2.0.
Installing package(s) ‘plyr’
trying URL 'http://cran.rstudio.com/bin/windows/contrib/3.2/plyr_1.8.2.zip'
Content type 'application/zip' length 1128621 bytes (1.1 MB)
downloaded 1.1 MB
package ‘plyr’ successfully unpacked and MD5 sums checked
The downloaded binary packages are in
C:\Users\brb\AppData\Local\Temp\RtmpyUjsJD\downloaded_packages
> library(lumi)
Error in loadNamespace(j <- i[[1L]], c(lib.loc, .libPaths()), versionCheck = vI[[j]]) :
there is no package called ‘mclust’
Error: package or namespace load failed for ‘lumi’
> ?biocLite
Warning messages:
1: In read.dcf(file.path(p, "DESCRIPTION"), c("Package", "Version")) :
cannot open compressed file 'C:/Users/brb/Documents/R/win-library/3.2/Biostrings/DESCRIPTION', probable reason 'No such file or directory'
2: In find.package(if (is.null(package)) loadedNamespaces() else package, :
there is no package called ‘Biostrings’
> library(lumi)
Error in loadNamespace(j <- i[[1L]], c(lib.loc, .libPaths()), versionCheck = vI[[j]]) :
there is no package called ‘mclust’
In addition: Warning messages:
1: In read.dcf(file.path(p, "DESCRIPTION"), c("Package", "Version")) :
cannot open compressed file 'C:/Users/brb/Documents/R/win-library/3.2/Biostrings/DESCRIPTION', probable reason 'No such file or directory'
2: In find.package(if (is.null(package)) loadedNamespaces() else package, :
there is no package called ‘Biostrings’
Error: package or namespace load failed for ‘lumi’
Other people also have the similar problem. The possible cause is the virus scanner locks the file and R cannot move them.
Some possible solutions:
- Delete ALL folders under R/library (e.g. C:/Progra~1/R/R-3.2.0/library) folder and install the main package again using install.packages() or biocLite().
- For specific package like 'lumi' from Bioconductor, we can find out all dependency packages and then install them one by one.
- Find out and install the top level package which misses dependency packages.
- This is based on the fact that install.packages() or biocLite() sometimes just checks & installs the 'Depends' and 'Imports' packages and won't install all packages recursively
- we can do a small experiment by removing a package which is not directly dependent/imported by another package; e.g. 'iterators' is not dependent/imported by 'glment' directly but indirectly. So if we run remove.packages("iterators"); install.packages("glmnet"), then the 'iterator' package is still missing.
- A real example is if the missing packages are 'Biostrings', 'limma', 'mclust' (these are packages that 'minfi' directly depends/imports although they should be installed when I run biocLite("lumi") command), then I should just run the command remove.packages("minfi"); biocLite("minfi"). If we just run biocLite("lumi") or biocLite("methylumi"), the missing packages won't be installed.
Error in download.file(url, destfile, method, mode = "wb", ...)
HTTP status was '404 Not Found'
Tested on an existing R-3.2.0 session. Note that VariantAnnotation 1.14.4 was just uploaded to Bioc.
> biocLite("COSMIC.67")
BioC_mirror: http://bioconductor.org
Using Bioconductor version 3.1 (BiocInstaller 1.18.3), R version 3.2.0.
Installing package(s) ‘COSMIC.67’
also installing the dependency ‘VariantAnnotation’
trying URL 'http://bioconductor.org/packages/3.1/bioc/bin/windows/contrib/3.2/VariantAnnotation_1.14.3.zip'
Error in download.file(url, destfile, method, mode = "wb", ...) :
cannot open URL 'http://bioconductor.org/packages/3.1/bioc/bin/windows/contrib/3.2/VariantAnnotation_1.14.3.zip'
In addition: Warning message:
In download.file(url, destfile, method, mode = "wb", ...) :
cannot open: HTTP status was '404 Not Found'
Warning in download.packages(pkgs, destdir = tmpd, available = available, :
download of package ‘VariantAnnotation’ failed
installing the source package ‘COSMIC.67’
trying URL 'http://bioconductor.org/packages/3.1/data/experiment/src/contrib/COSMIC.67_1.4.0.tar.gz'
Content type 'application/x-gzip' length 40999037 bytes (39.1 MB)
However, when I tested on a new R-3.2.0 (just installed in VM), the COSMIC package installation is successful. That VariantAnnotation version 1.14.4 was installed (this version was just updated today from Bioconductor).
The cause of the error is the available.package() function will read the rds file first from cache in a tempdir (C:\Users\XXXX\AppData\Local\Temp\RtmpYYYYYY). See lines 51-55 of <packages.R>.
dest <- file.path(tempdir(),
paste0("repos_", URLencode(repos, TRUE), ".rds"))
if(file.exists(dest)) {
res0 <- readRDS(dest)
} else {
...
}
Since my R was opened 1 week ago, the rds file it reads today contains old information. Note that Bioconductor does not hold the source code or binary code for the old version of packages. This explains why biocLite() function broke. When I restart R, the original problem is gone.
If we look at the source code of available.packages(), we will see we could use cacheOK option in download.file() function.
download.file(url, destfile, method, cacheOK = FALSE, quiet = TRUE, mode ="wb")
Another case: Error in download.file(url, destfile, method, mode = "wb", ...)
> install.packages("quantreg")
There is a binary version available but the source version is later:
binary source needs_compilation
quantreg 5.33 5.34 TRUE
Do you want to install from sources the package which needs compilation?
y/n: n
trying URL 'https://cran.rstudio.com/bin/macosx/el-capitan/contrib/3.4/quantreg_5.33.tgz'
Warning in install.packages :
cannot open URL 'https://cran.rstudio.com/bin/macosx/el-capitan/contrib/3.4/quantreg_5.33.tgz': HTTP status was '404 Not Found'
Error in download.file(url, destfile, method, mode = "wb", ...) :
cannot open URL 'https://cran.rstudio.com/bin/macosx/el-capitan/contrib/3.4/quantreg_5.33.tgz'
Warning in install.packages :
download of package ‘quantreg’ failed
It seems the binary package cannot be found on the mirror. So the solution here is to download the package from the R main server. Note that after I have successfully installed the binary package from the main R server, I remove the package in R and try to install the binary package from rstudio.com server agin and it works this time.
> install.packages("quantreg", repos = "https://cran.r-project.org")
trying URL 'https://cran.r-project.org/bin/macosx/el-capitan/contrib/3.4/quantreg_5.34.tgz'
Content type 'application/x-gzip' length 1863561 bytes (1.8 MB)
==================================================
downloaded 1.8 MB
Another case: Error in download.file() on Windows 7
For some reason, IE 8 cannot interpret https://ftp.ncbi.nlm.nih.gov though it understands ftp://ftp.ncbi.nlm.nih.gov.
This is tested using R 3.4.3.
> download.file("https://ftp.ncbi.nlm.nih.gov/geo/series/GSE7nnn/GSE7848/soft/GSE7848_family.soft.gz", "test.soft.gz")
trying URL 'https://ftp.ncbi.nlm.nih.gov/geo/series/GSE7nnn/GSE7848/soft/GSE7848_family.soft.gz'
Error in download.file("https://ftp.ncbi.nlm.nih.gov/geo/series/GSE7nnn/GSE7848/soft/GSE7848_family.soft.gz", :
cannot open URL 'https://ftp.ncbi.nlm.nih.gov/geo/series/GSE7nnn/GSE7848/soft/GSE7848_family.soft.gz'
In addition: Warning message:
In download.file("https://ftp.ncbi.nlm.nih.gov/geo/series/GSE7nnn/GSE7848/soft/GSE7848_family.soft.gz", :
InternetOpenUrl failed: 'An error occurred in the secure channel support'
> download.file("ftp://ftp.ncbi.nlm.nih.gov/geo/series/GSE7nnn/GSE7848/soft/GSE7848_family.soft.gz", "test.soft.gz")
trying URL 'ftp://ftp.ncbi.nlm.nih.gov/geo/series/GSE7nnn/GSE7848/soft/GSE7848_family.soft.gz'
downloaded 9.1 MB
ERROR: failed to lock directory
Follow the suggestion to remove the LOCK file. See the post.
It could happened in calling install.packages(), biocLite() or devtools::install_github(), and so on.
Error in unloadNamespace(package)
> d3heatmap(mtcars, scale = "column", colors = "Blues")
Error: 'col_numeric' is not an exported object from 'namespace:scales'
> packageVersion("scales")
[1] ‘0.2.5’
> library(scales)
Error in unloadNamespace(package) :
namespace ‘scales’ is imported by ‘ggplot2’ so cannot be unloaded
In addition: Warning message:
package ‘scales’ was built under R version 3.2.1
Error in library(scales) :
Package ‘scales’ version 0.2.4 cannot be unloaded
> search()
[1] ".GlobalEnv" "package:d3heatmap" "package:ggplot2"
[4] "package:microbenchmark" "package:COSMIC.67" "package:BiocInstaller"
[7] "package:stats" "package:graphics" "package:grDevices"
[10] "package:utils" "package:datasets" "package:methods"
[13] "Autoloads" "package:base"
If I open a new R session, the above error will not happen!
The problem occurred because the 'scales' package version required by the d3heatmap package/function is old. See this post. And when I upgraded the 'scales' package, it was locked by the package was imported by the ggplot2 package.
Unload a package
See an example below.
require(splines) detach(package:splines, unload=TRUE)
New R packages as reported by CRANberries
http://blog.revolutionanalytics.com/2015/07/mranspackages-spotlight.html
#----------------------------
# SCRAPE CRANBERRIES FILES TO COUNT NEW PACKAGES AND PLOT
#
library(ggplot2)
# Build a vextor of the directories of interest
year <- c("2013","2014","2015")
month <- c("01","02","03","04","05","06","07","08","09","10","11","12")
span <-c(rep(month,2),month[1:7])
dir <- "http://dirk.eddelbuettel.com/cranberries"
url2013 <- file.path(dir,"2013",month)
url2014 <- file.path(dir,"2014",month)
url2015 <- file.path(dir,"2015",month[1:7])
url <- c(url2013,url2014,url2015)
# Read each directory and count the new packages
new_p <- vector()
for(i in url){
raw.data <- readLines(i)
new_p[i] <- length(grep("New package",raw.data,value=TRUE))
}
# Plot
time <- seq(as.Date("2013-01-01"), as.Date("2015-07-01"), by="months")
new_pkgs <- data.frame(time,new_p)
ggplot(new_pkgs, aes(time,y=new_p)) +
geom_line() + xlab("") + ylab("Number of new packages") +
geom_smooth(method='lm') + ggtitle("New R packages as reported by CRANberries")
Top new packages in 2015
- 2015 R packages roundup by CHRISTOPH SAFFERLING
- R trends in 2015 by MAX GORDON
Speeding up package installation
- http://blog.jumpingrivers.com/posts/2017/speed_package_installation/
- (Much) Faster Package (Re-)Installation via Caching
- (Much) Faster Package (Re-)Installation via Caching, part 2
package ‘XXX’ was installed by an R version with different internals
it needs to be reinstalled for use with this R version. The problem seems to be specific to R 3.5.1 in Ubuntu 16.04. I got this message when I try to install the keras and tidyverse packages. The 'XXX' package includes nlme for installing "tidyverse" and Matrix for installing "reticulate". I have already logged in as root. I need to manually install these packages again though it seems I did not see a version change for these packages.
Today it also happened when I tried to install "pec" which broke when it was installing "Hmisc". The error message is "Error : package ‘rpart’ was installed by an R version with different internals; it needs to be reinstalled for use with this R version". I am using R 3.5.2. rpart version is ‘4.1.13’. The solution is I install rpart again (under my account is enough) though rpart does not have a new version. Then I can install "Hmisc".
packrat (reproducible search): project specific package managment
- CRAN & Github
- Videos:
- Packrat will not only store all packages, but also all project files.
- Packrat is integrated in RStudio’s user interface. It allows you to share projects along co-workers easily. See Using Packrat with RStudio.
- limitations.
- Git and packrat. The packrat/src directory can be very large. If you don't want them available in your git-repo, you simply add packrat/src/ to the .gitignore. But, this will mean that anyone accessing the git-repo will not have access to the package source code, and the files will be downloaded from CRAN, or from wherever the source line dictates within the packrat.lock file.
- A scenario that we need packrat: suppose we are developing a package in the current R-3.5.X. Our package requires the 'doRNG' package. That package depends the 'rngtools' package. A few months later a new R (3.6.0) was released and a new release (1.3.1.1) of 'rngtools' also requires R-3.6.0. So if we want to install 'doRNG' in R-3.5.x, it will fail with an error: dependency 'rngtools' is not available for package 'doRNG' .
Create a snapshot
- Do we really need to call packrat::snapshot()? The walk through page says it is not needed but the lock file is not updated from my testing.
- I got an error when it is trying to fetch the source code from bioconductor and local repositories: packrat is trying to fetch the source from CRAN in these two packages.
- On normal case, the packrat/packrat.lock file contains two entries in 'Repos' field (line 4).
- The cause of the error is I ran snapshot() after I quitted R and entered again. So the solution is to add bioc and local repositories to options(repos).
- So what is important of running snapshot()?
- Check out the forum.
> dir.create("~/projects/babynames", recu=T)
> packrat::init("~/projects/babynames")
Initializing packrat project in directory:
- "~/projects/babynames"
Adding these packages to packrat:
_
packrat 0.4.9-3
Fetching sources for packrat (0.4.9-3) ... OK (CRAN current)
Snapshot written to '/home/brb/projects/babynames/packrat/packrat.lock'
Installing packrat (0.4.9-3) ...
OK (built source)
Initialization complete!
Unloading packages in user library:
- packrat
Packrat mode on. Using library in directory:
- "~/projects/babynames/packrat/lib"
> install.packages("reshape2")
> packrat::snapshot()
> system("tree -L 2 ~/projects/babynames/packrat/")
/home/brb/projects/babynames/packrat/
├── init.R
├── lib
│ └── x86_64-pc-linux-gnu
├── lib-ext
│ └── x86_64-pc-linux-gnu
├── lib-R # base packages
│ └── x86_64-pc-linux-gnu
├── packrat.lock
├── packrat.opts
└── src
├── bitops
├── glue
├── magrittr
├── packrat
├── plyr
├── Rcpp
├── reshape2
├── stringi
└── stringr
Restoring snapshots
Suppose a packrat project was created on Ubuntu 16.04 and we now want to repeat the analysis on Ubuntu 18.04. We first copy the whole project directory ('babynames') to Ubuntu 18.04. Then we should delete the library subdirectory ('packrat/lib') which contains binary files (*.so) that do not work on the new OS. After we delete the library subdirectory, start R from the project directory. Now if we run packrat::restore() command, it will re-install all missing libraries. Bingo! NOTE: Maybe I should use packrat::bundle() instead of manually copy the whole project folder.
Note: some OS level libraries (e.g. libXXX-dev) need to be installed manually beforehand in order for the magic to work.
$ rm -rf ~/projects/babynames/packrat/lib
$ cd ~/projects/babynames/
$ R
>
> packrat::status()
> remove.packages("plyr")
> packrat::status()
> packrat::restore()
Set Up a Custom CRAN-like Repository
See https://rstudio.github.io/packrat/custom-repos.html. Note the personal repository name ('sushi' in this example) used in "Repository" field of the personal package will be used in <packrat/packrat.lock> file. So as long as we work on the same computer, it is easy to restore a packrat project containing packages coming from personal repository.
- packrat::init()
- packrat::snapshot(), packrat::restore()
- packrat::clean()
- packrat::status()
- packrat::install_local() # http://rstudio.github.io/packrat/limitations.html
- packrat::bundle() # see @28:44 of the video, packrat::unbundle() # see @29:17 of the same video. This will rebuild all packages
- packrat::on(), packrat::off()
- packrat::get_opts()
- packrat::set_opts() # http://rstudio.github.io/packrat/limitations.html
- packrat::opts$local.repos("~/local-cran")
- packrat::opts$external.packages(c("devtools")) # break the isolation
- packrat::extlib()
- packrat::with_extlib()
- packrat::project_dir(), .libPaths()
Warning
- If we download and modify some function definition from a package in CRAN without changing DESCRIPTION file or the package name, the snapshot created using packrat::snapshot() will contain the package source from CRAN instead of local repository. This is because (I guess) the DESCRIPTION file contains a field 'Repository' with the value 'CRAN'.
Docker
Docker and Packrat. The first part of generating packrat.lock is not quite right since the file was generated in the container only. We should use -v in the docker run command. The github repository at https://github.com/joelnitta/docker-packrat-example has fixed the problem.
$ git clone https://github.com/joelnitta/docker-packrat-example.git
$ cd docker-packrat-example
# Step 1: create the 'packrat.lock' file
$ nano install_packages.R # note: nano is not available in the rstudio container
# need to install additional OS level packages like libcurl
# in rocker/rstudio. Probably rocker/tidyverse is better than rstudio
#
$ docker run -it -e DISABLE_AUTH=true -v $(pwd):/home/rstudio/project rocker/tidyverse:3.6.0 bash
# Inside the container now
$ cd home/rstudio/project
$ time Rscript install_packages.R # generate 'packrat/packrat.lock'
$ exit # It took 43 minutes.
# Question: is there an easier way to generate packrat.lock without
# wasting time to install lots of packages?
# Step 2: build the image
# Open another terminal/tab
$ nano Dockerfile # change rocker image and R version. Make sure these two are the same as
# we have used when we created the 'packrat.lock' file
$ time docker build . -t mycontainer # It took 45 minutes.
$ docker run -it mycontainer R
# Step 3: check the packages defined in 'install_packages.R' are installed
packageVersion("minimal")
packageVersion("biospear")
Questions:
- After running the statement packrat::init(), it will leave a footprint of a hidden file ".Rprofile" in the current directory.
#### -- Packrat Autoloader (version 0.5.0) -- #### source("packrat/init.R") #### -- End Packrat Autoloader -- ####
- If the 'packrat' directory was accidentally deleted, next time when you launch R it will show an error message because it cannot find the file.
- The ownership of the 'packrat' directory will be root now. See this Package Management for Reproducible R Code.
- This sophisticated approach does not save the package source code. If a package has been updated and the version we used has been moved to archive in CRAN, what will happen when we try to restore it?
renv: successor to the packrat package
https://rstudio.github.io/renv/index.html
R package dependencies
Depends, Imports, Suggests, Enhances, LinkingTo
See Writing R Extensions and install.packages().
- Depends: list of package names which this package depends on. Those packages will be attached (so it is better to use Imports instead of Depends as much as you can) before the current package when library or require is called. The ‘Depends’ field can also specify a dependence on a certain version of R.
- Imports: lists packages whose namespaces are imported from (as specified in the NAMESPACE file) but which do not need to be attached.
- Suggests: lists packages that are not necessarily needed. This includes packages used only in examples, tests or vignettes, and packages loaded in the body of functions.
- Enhances: lists packages “enhanced” by the package at hand, e.g., by providing methods for classes from these packages, or ways to handle objects from these packages.
- LinkingTo: A package that wishes to make use of header files in other packages needs to declare them as a comma-separated list in the field ‘LinkingTo’ in the DESCRIPTION file.
- available.packages(); see packageStatus().
- download.packages()
- packageStatus(), update(), upgrade(). packageStatus() will return a list with two components:
- inst - a data frame with columns as the matrix returned by installed.packages plus "Status", a factor with levels c("ok", "upgrade"). Note: the manual does not mention "unavailable" case (but I do get it) in R 3.2.0?
- avail - a data frame with columns as the matrix returned by available.packages plus "Status", a factor with levels c("installed", "not installed", "unavailable"). Note: I don't get the "unavailable" case in R 3.2.0?
> x <- packageStatus()
> names(x)
[1] "inst" "avail"
> dim(x[['inst']])
[1] 225 17
> x[['inst']][1:3, ]
Package LibPath Version Priority Depends Imports
acepack acepack C:/Program Files/R/R-3.1.2/library 1.3-3.3 <NA> <NA> <NA>
adabag adabag C:/Program Files/R/R-3.1.2/library 4.0 <NA> rpart, mlbench, caret <NA>
affxparser affxparser C:/Program Files/R/R-3.1.2/library 1.38.0 <NA> R (>= 2.6.0) <NA>
LinkingTo Suggests Enhances
acepack <NA> <NA> <NA>
adabag <NA> <NA> <NA>
affxparser <NA> R.oo (>= 1.18.0), R.utils (>= 1.32.4),\nAffymetrixDataTestFiles <NA>
License License_is_FOSS License_restricts_use OS_type MD5sum NeedsCompilation Built
acepack MIT + file LICENSE <NA> <NA> <NA> <NA> yes 3.1.2
adabag GPL (>= 2) <NA> <NA> <NA> <NA> no 3.1.2
affxparser LGPL (>= 2) <NA> <NA> <NA> <NA> <NA> 3.1.1
Status
acepack ok
adabag ok
affxparser unavailable
> dim(x[['avail']])
[1] 6538 18
> x[['avail']][1:3, ]
Package Version Priority Depends Imports LinkingTo
A3 A3 0.9.2 <NA> R (>= 2.15.0), xtable, pbapply <NA> <NA>
ABCExtremes ABCExtremes 1.0 <NA> SpatialExtremes, combinat <NA> <NA>
ABCanalysis ABCanalysis 1.0.1 <NA> R (>= 2.10) Hmisc, plotrix <NA>
Suggests Enhances License License_is_FOSS License_restricts_use OS_type Archs
A3 randomForest, e1071 <NA> GPL (>= 2) <NA> <NA> <NA> <NA>
ABCExtremes <NA> <NA> GPL-2 <NA> <NA> <NA> <NA>
ABCanalysis <NA> <NA> GPL-3 <NA> <NA> <NA> <NA>
MD5sum NeedsCompilation File Repository Status
A3 <NA> <NA> <NA> http://cran.rstudio.com/bin/windows/contrib/3.1 not installed
ABCExtremes <NA> <NA> <NA> http://cran.rstudio.com/bin/windows/contrib/3.1 not installed
ABCanalysis <NA> <NA> <NA> http://cran.rstudio.com/bin/windows/contrib/3.1 not installed
- packageVersion(), packageDescription()
- install.packages(), remove.packages().
- installed.packages(); see packageStatus().
- update.packages(), old.packages(), new.packages()
- setRepositories()
- contrib.url()
- chooseCRANmirror(), chooseBioCmirror()
- suppressForeignCheck()
tools package
- https://www.rdocumentation.org/packages/tools/versions/3.6.1
- CRAN_package_db() from ?CRANtools
- dependsOnPkgs()
- package.dependencies(), pkgDepends(), etc are deprecated now, mostly in favor of package_dependencies() which is both more flexible and efficient. See R 3.3.0 News.
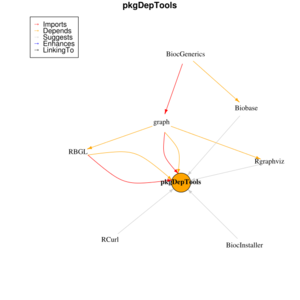
Bioconductor's pkgDepTools package
The is an example of querying the dependencies of the notorious 'lumi' package which often broke the installation script. I am using R 3.2.0 and Bioconductor 3.1.
The getInstallOrder function is useful to get a list of all (recursive) dependency packages.
source("http://bioconductor.org/biocLite.R")
if (!require(pkgDepTools)) {
biocLite("pkgDepTools", ask = FALSE)
library(pkgDepTools)
}
MkPlot <- FALSE
library(BiocInstaller)
biocUrl <- biocinstallRepos()["BioCsoft"]
biocDeps <- makeDepGraph(biocUrl, type="source", dosize=FALSE) # pkgDepTools defines its makeDepGraph()
PKG <- "lumi"
if (MkPlot) {
if (!require(Biobase)) {
biocLite("Biobase", ask = FALSE)
library(Biobase)
}
if (!require(Rgraphviz)) {
biocLite("Rgraphviz", ask = FALSE)
library(Rgraphviz)
}
categoryNodes <- c(PKG, names(acc(biocDeps, PKG)[[1]]))
categoryGraph <- subGraph(categoryNodes, biocDeps)
nn <- makeNodeAttrs(categoryGraph, shape="ellipse")
plot(categoryGraph, nodeAttrs=nn) # Complete but plot is too complicated & font is too small.
}
system.time(allDeps <- makeDepGraph(biocinstallRepos(), type="source",
keep.builtin=TRUE, dosize=FALSE)) # takes a little while
# user system elapsed
# 175.737 10.994 186.875
# Warning messages:
# 1: In .local(from, to, graph) : edges replaced: ‘SNPRelate|gdsfmt’
# 2: In .local(from, to, graph) :
# edges replaced: ‘RCurl|methods’, ‘NA|bitops’
# When needed.only=TRUE, only those dependencies not currently installed are included in the list.
x1 <- sort(getInstallOrder(PKG, allDeps, needed.only=TRUE)$packages); x1
[1] "affy" "affyio"
[3] "annotate" "AnnotationDbi"
[5] "base64" "beanplot"
[7] "Biobase" "BiocParallel"
[9] "biomaRt" "Biostrings"
[11] "bitops" "bumphunter"
[13] "colorspace" "DBI"
[15] "dichromat" "digest"
[17] "doRNG" "FDb.InfiniumMethylation.hg19"
[19] "foreach" "futile.logger"
[21] "futile.options" "genefilter"
[23] "GenomeInfoDb" "GenomicAlignments"
[25] "GenomicFeatures" "GenomicRanges"
[27] "GEOquery" "ggplot2"
[29] "gtable" "illuminaio"
[31] "IRanges" "iterators"
[33] "labeling" "lambda.r"
[35] "limma" "locfit"
[37] "lumi" "magrittr"
[39] "matrixStats" "mclust"
[41] "methylumi" "minfi"
[43] "multtest" "munsell"
[45] "nleqslv" "nor1mix"
[47] "org.Hs.eg.db" "pkgmaker"
[49] "plyr" "preprocessCore"
[51] "proto" "quadprog"
[53] "RColorBrewer" "Rcpp"
[55] "RCurl" "registry"
[57] "reshape" "reshape2"
[59] "rngtools" "Rsamtools"
[61] "RSQLite" "rtracklayer"
[63] "S4Vectors" "scales"
[65] "siggenes" "snow"
[67] "stringi" "stringr"
[69] "TxDb.Hsapiens.UCSC.hg19.knownGene" "XML"
[71] "xtable" "XVector"
[73] "zlibbioc"
# When needed.only=FALSE the complete list of dependencies is given regardless of the set of currently installed packages.
x2 <- sort(getInstallOrder(PKG, allDeps, needed.only=FALSE)$packages); x2
[1] "affy" "affyio" "annotate"
[4] "AnnotationDbi" "base64" "beanplot"
[7] "Biobase" "BiocGenerics" "BiocInstaller"
[10] "BiocParallel" "biomaRt" "Biostrings"
[13] "bitops" "bumphunter" "codetools"
[16] "colorspace" "DBI" "dichromat"
[19] "digest" "doRNG" "FDb.InfiniumMethylation.hg19"
[22] "foreach" "futile.logger" "futile.options"
[25] "genefilter" "GenomeInfoDb" "GenomicAlignments"
[28] "GenomicFeatures" "GenomicRanges" "GEOquery"
[31] "ggplot2" "graphics" "grDevices"
[34] "grid" "gtable" "illuminaio"
[37] "IRanges" "iterators" "KernSmooth"
[40] "labeling" "lambda.r" "lattice"
[43] "limma" "locfit" "lumi"
[46] "magrittr" "MASS" "Matrix"
[49] "matrixStats" "mclust" "methods"
[52] "methylumi" "mgcv" "minfi"
[55] "multtest" "munsell" "nleqslv"
[58] "nlme" "nor1mix" "org.Hs.eg.db"
[61] "parallel" "pkgmaker" "plyr"
[64] "preprocessCore" "proto" "quadprog"
[67] "RColorBrewer" "Rcpp" "RCurl"
[70] "registry" "reshape" "reshape2"
[73] "rngtools" "Rsamtools" "RSQLite"
[76] "rtracklayer" "S4Vectors" "scales"
[79] "siggenes" "snow" "splines"
[82] "stats" "stats4" "stringi"
[85] "stringr" "survival" "tools"
[88] "TxDb.Hsapiens.UCSC.hg19.knownGene" "utils" "XML"
[91] "xtable" "XVector" "zlibbioc"
> sort(setdiff(x2, x1)) # Not all R's base packages are included; e.g. 'base', 'boot', ...
[1] "BiocGenerics" "BiocInstaller" "codetools" "graphics" "grDevices"
[6] "grid" "KernSmooth" "lattice" "MASS" "Matrix"
[11] "methods" "mgcv" "nlme" "parallel" "splines"
[16] "stats" "stats4" "survival" "tools" "utils"
Bioconductor BiocPkgTools
Collection of simple tools for learning about Bioc Packages. Functionality includes access to :
- Download statistics
- General package listing
- Build reports
- Package dependency graphs
- Vignettes
Overview of BiocPkgTools & Dependency graphs
BiocPkgTools: Toolkit for Mining the Bioconductor Package Ecosystem in biorxiv.org.
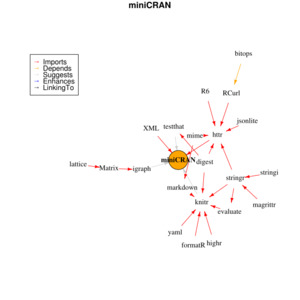
miniCRAN package
miniCRAN package can be used to identify package dependencies or create a local CRAN repository. It can be used on repositories other than CRAN, such as Bioconductor.
- http://blog.revolutionanalytics.com/2014/07/dependencies-of-popular-r-packages.html
- http://www.r-bloggers.com/introducing-minicran-an-r-package-to-create-a-private-cran-repository/
- http://www.magesblog.com/2014/09/managing-r-package-dependencies.html
- Using miniCRAN in Azure ML
- developing internal CRAN Repositories
Before we go into R, we need to install some packages from Ubuntu terminal. See here.
# Consider glmnet package (today is 4/29/2015)
# Version: 2.0-2
# Depends: Matrix (≥ 1.0-6), utils, foreach
# Suggests: survival, knitr, lars
if (!require("miniCRAN")) {
install.packages("miniCRAN", dependencies = TRUE, repos="http://cran.rstudio.com") # include 'igraph' in Suggests.
library(miniCRAN)
}
if (!"igraph" %in% installed.packages()[,1]) install.packages("igraph")
tags <- "glmnet"
pkgDep(tags, suggests=TRUE, enhances=TRUE) # same as pkgDep(tags)
# [1] "glmnet" "Matrix" "foreach" "codetools" "iterators" "lattice" "evaluate" "digest"
# [9] "formatR" "highr" "markdown" "stringr" "yaml" "mime" "survival" "knitr"
# [17] "lars"
dg <- makeDepGraph(tags, suggests=TRUE, enhances=TRUE) # miniCRAN defines its makeDepGraph()
plot(dg, legendPosition = c(-1, 1), vertex.size=20)
We can also display the dependence for a package from the Bioconductor repository.
tags <- "DESeq2"
# Depends S4Vectors, IRanges, GenomicRanges, Rcpp (>= 0.10.1), RcppArmadillo (>= 0.3.4.4)
# Imports BiocGenerics(>= 0.7.5), Biobase, BiocParallel, genefilter, methods, locfit, geneplotter, ggplot2, Hmisc
# Suggests RUnit, gplots, knitr, RColorBrewer, BiocStyle, airway,\npasilla (>= 0.2.10), DESeq, vsn
# LinkingTo Rcpp, RcppArmadillo
index <- function(url, type="source", filters=NULL, head=5, cols=c("Package", "Version")){
contribUrl <- contrib.url(url, type=type)
available.packages(contribUrl, type=type, filters=filters)
}
bioc <- local({
env <- new.env()
on.exit(rm(env))
evalq(source("http://bioconductor.org/biocLite.R", local=TRUE), env)
biocinstallRepos() # return URLs
})
bioc
# BioCsoft
# "http://bioconductor.org/packages/3.0/bioc"
# BioCann
# "http://bioconductor.org/packages/3.0/data/annotation"
# BioCexp
# "http://bioconductor.org/packages/3.0/data/experiment"
# BioCextra
# "http://bioconductor.org/packages/3.0/extra"
# CRAN
# "http://cran.fhcrc.org"
# CRANextra
# "http://www.stats.ox.ac.uk/pub/RWin"
str(index(bioc["BioCsoft"])) # similar to cranJuly2014 object
system.time(dg <- makeDepGraph(tags, suggests=TRUE, enhances=TRUE, availPkgs = index(bioc["BioCsoft"]))) # Very quick!
plot(dg, legendPosition = c(-1, 1), vertex.size=20)
The dependencies of GenomicFeature and GenomicAlignments are more complicated. So we turn the 'suggests' option to FALSE.
tags <- "GenomicAlignments" dg <- makeDepGraph(tags, suggests=FALSE, enhances=FALSE, availPkgs = index(bioc["BioCsoft"])) plot(dg, legendPosition = c(-1, 1), vertex.size=20)
MRAN (CRAN only)
cranly
Reverse dependence
- http://romainfrancois.blog.free.fr/index.php?post/2011/10/30/Rcpp-reverse-dependency-graph
- Checking reverse dependencies: the tiny way
Install packages offline
http://www.mango-solutions.com/wp/2017/05/installing-packages-without-internet/
Install a packages locally and its dependencies
It's impossible to install the dependencies if you want to install a package locally. See Windows-GUI: "Install Packages from local zip files" and dependencies
A minimal R package (for testing purpose)
https://github.com/joelnitta/minimal. Question: is there a one from CRAN?
An R package that does not require others during install.packages()
Create a new R package, namespace, documentation
- http://cran.r-project.org/doc/contrib/Leisch-CreatingPackages.pdf (highly recommend)
- https://stat.ethz.ch/pipermail/r-devel/2013-July/066975.html
- Benefit of import in a namespace
- This youtube video from Tyler Rinker teaches how to use RStudio to develop an R package and also use Git to do version control. Very useful!
- Developing R packages by Jeff Leek in Johns Hopkins University.
- R packages book by Hadley Wickham.
- R package primer a minimal tutorial from Karl Broman.
- How to make and share an R package in 3 steps (6/14/2017)
- Package Development tutorial for useR! 2019 Toulouse
Install software for PDF output
- Windows: Miktex and pandoc (Create HTML or PDF Files with R, Knitr, MiKTeX, and Pandoc)
- Unix: TeX Live (sudo apt-get install texlive-full) and pandoc
- Mac: MacTex
Windows: Rtools
Installing RTools for Compiled Code via Rcpp. Just remember to check the option to include some paths in the PATH environment variable.
R CMD
R Installation and Administration
- R CMD build XXX. Note this will not create pdf files for vignettes. The output is a tarball.
- R CMD INSTALL
- R CMD REMOVE
- R CMD SHLIB files. For example, "Rcmd shlib *.f *.c -o surv.dll" on Windows.
- R CMD make
- R CMD check XXX. Useful if we want to create reference manual (PDF file). See R create reference manual with R CMD check.
- R CMD javareconf
usethis package
- https://usethis.r-lib.org/. usethis is a workflow package: it automates repetitive tasks that arise during project setup and development, both for R packages and non-package projects.
- usethis 1.5.0
R package depends vs imports
- http://stackoverflow.com/questions/8637993/better-explanation-of-when-to-use-imports-depends
- http://stackoverflow.com/questions/9893791/imports-and-depends
- https://stat.ethz.ch/pipermail/r-devel/2013-August/067082.html
In the namespace era Depends is never really needed. All modern packages have no technical need for Depends anymore. Loosely speaking the only purpose of Depends today is to expose other package's functions to the user without re-exporting them.
load = functions exported in myPkg are available to interested parties as myPkg::foo or via direct imports - essentially this means the package can now be used
attach = the namespace (and thus all exported functions) is attached to the search path - the only effect is that you have now added the exported functions to the global pool of functions - sort of like dumping them in the workspace (for all practical purposes, not technically)
import a function into a package = make sure that this function works in my package regardless of the search path (so I can write fn1 instead of pkg1::fn1 and still know it will come from pkg1 and not someone's workspace or other package that chose the same name)
The distinction is between "loading" and "attaching" a package. Loading it (which would be done if you had MASS::loglm, or imported it) guarantees that the package is initialized and in memory, but doesn't make it visible to the user without the explicit MASS:: prefix. Attaching it first loads it, then modifies the user's search list so the user can see it.
Loading is less intrusive, so it's preferred over attaching. Both library() and require() would attach it.
R package suggests
stringr has suggested htmlwidgets. An error will come out if the suggested packages are not available.
> library(stringr)
> str_view(c("abc", "a.c", "bef"), "a\\.c")
Error in loadNamespace(name) : there is no package called ‘htmlwidgets’
Useful functions for accessing files in packages
- system.file()
- path.package() and normalizePath().
> system.file(package = "batr")
[1] "f:/batr"
> system.file("extdata", package = "batr")
> path.package("batr")
[1] "f:\\batr"
# sometimes it returns the forward slash format for some reason; C:/Program Files/R/R-3.4.0/library/batr
# so it is best to add normalizePath().
> normalizePath(path.package("batr"))
RStudio shortcuts
Create R package with devtools and roxygen2
- A useful post by Jacob Montgomery. Watch the youtube video there. The process requires 3 components: RStudio software, devtools and roxygen2 (creating documentation from R code) packages.
- MAKING PACKAGES IN R USING DEVTOOLS
- R code workflow from Hadley Wickham.
- RStudio:addins part 2 - roxygen documentation formatting made easy
- Instructions for Creating Your Own R Package. It includes creating a R package with functions written in C++ via Rcpp helper function.
- devtools cheatsheet (2 pages)
How to use devtools::load_all("FolderName"). load_all() loads any modified R files, and recompile and reload any modified C or Fortran files.
# Step 1
library(devtools)
# Step 2
dir.create(file.path("MyCode", "R"), recursive = TRUE)
cat("foo=function(x){x*2}", file = file.path("MyCode", "R", "foo.R"))
write.dcf(list(Package = "MyCode", Title = "My Code for this project", Description = "To tackle this problem",
Version = "0.0", License = "For my eyes only", Author = "First Last <[email protected]>",
Maintainer = "First Last <[email protected]>"), file = file.path("MyCode", "DESCRIPTION"))
# OR
# create("path/to/package/pkgname")
# create() will create R/ directory, DESCRIPTION and NAMESPACE files.
# Step 3 (C/Fortran code, optional)
dir.create(file.path("MyCode", "src"))
cat("void cfoo(double *a, double *b, double *c){*c=*a+*b;}\n", file = file.path("MyCode",
"src", "cfoo.c"))
cat("useDynLib(MyCode)\n", file = file.path("MyCode", "NAMESPACE"))
# Step 4
load_all("MyCode")
# Step 5
# Modify R/C/Fortran code and run load_all("MyCode")
# Step 6 (Automatically generate the documentation, optional)
document()
# Step 7 (Deployment, optional)
build("MyCode")
# Step 8 (Install the package, optional)
install()
Note:
- load_all("FolderName") will make the FolderName to become like a package to be loaded into the current R session so the 2nd item returned from search() will be "package:FolderName". However, the FolderName does not exist under Program Files/R/R-X.Y.Z/library nor Documents/R/win-library/X.Y/ (Windows OS).
- build("FolderName") will create a tarball in the current directory. User can install the new package for example using Packages -> Install packages from local files on Windows OS.
- For the simplest R package, the source code only contains a file <DESCRIPTION> and a folder <R> with individual R files in the text format.
Create a package by RStudio
- How to Create and Distribute an R Package.
- The goal of the article to set up an easily installable R package on Github for others to use via remotes::install_github().
- The main tools required are RStudio along with the packages roxygen2 and usethis.
- How to take care of Bioconductor and Github Dependencies
- How to publish your package to Github
Binary packages
- No .R files in the R/ directory. There are 3 files that store the parsed functions in an efficient file format. This is the result of loading all the R code and then saving the functions with save().
- A Meta/ directory contains a number of Rds files. These files contain cached metadata about the package, like what topics the help files cover and parsed version of the DESCRIPTION file.
- An html/ directory.
- libs/ directory if you have any code in the src/' directory
- The contents of inst/ are moved to the top-level directory.
Vignette
- Add a static pdf vignette to an R package
- devtools::build_vignettes(). The files are copied in the 'doc' directory and an vignette index is created in 'Meta/vignette.rds', as they would be in a built package.
- build_vignette can't find functions
README.Rmd & README.md files
See Releasing a package from R packages by Hadley Wickham.
How to convert .Rmd into .md in R studio?
Example: ggplot2 repository at Github
It seems RStudio cannot create TOC for *.md files. Visual Studio Code has an extension to do that.
What is a library?
A library is simply a directory containing installed packages.
You can use .libPaths() to see which libraries are currently active.
.libPaths() lapply(.libPaths(), dir)
Object names
- Variable and function names should be lower case.
- Use an underscore (_) to separate words within a name (reserve . for S3 methods).
- Camel case is a legitimate alternative, but be consistent! For example, preProcess(), twoClassData, createDataPartition(), trainingRows, trainPredictors, testPredictors, trainClasses, testClasses have been used in Applied Predictive Modeling by Kuhn & Johnson.
- Generally, variable names should be nouns and function names should be verb.
Spacing
- Add a space around the operators +, -, \ and *.
- Include a space around the assignment operators, <- and =.
- Add a space around any comparison operators such as == and <.
Indentation
- Use two spaces to indent code.
- Never mix tabs and spaces.
- RStudio can automatically convert the tab character to spaces (see Tools -> Global options -> Code).
\dontrun{}
- Writing R Extensions > Documenting functions and search for dontrun
- How to not run an example using roxygen2?
- What does "not run" mean in R help pages? See ?example .
formatR package
Use formatR package to clean up poorly formatted code
install.packages("formatR")
formatR::tidy_dir("R")
Another way is to use the linter package.
install.packages("lintr")
lintr:::lin_package()
Minimal R package for submission
https://stat.ethz.ch/pipermail/r-devel/2013-August/067257.html and CRAN Repository Policy.
r-hub/rhub package: the R package builder service
https://github.com/r-hub/proposal
- rhub 1.1.1 is on CRAN! 2019/4/8
- R package developers, why should you care about R-hub? 2019/3/26
- Local Linux checks with Docker
- https://www.rstudio.com/resources/videos/r-hub-overview/
- http://blog.revolutionanalytics.com/2016/10/r-hub-public-beta.html
- Setting up continuous multi-platform R package building, checking and testing with R-Hub, Docker and GitLab CI/CD for free, with a working example 2019/4/27
- How to keep up with CRAN policies and processes? 2019/5/29
- Overview of the CRAN checks API 2019/6/10
Continuous Integration: Travis-CI (Linux, Mac)
- A Beginner's Guide to Travis-CI
- testhat package. Lots of packages suggest testhat; for example princurve.
- http://johnmuschelli.com/neuroc/getting_ready_for_submission/index.html#61_travis
Continuous Integration: Appveyor (Windows)
- Appveyor is a continuous integration service that builds projects on Windows machines.
- http://johnmuschelli.com/neuroc/getting_ready_for_submission/index.html#62_appveyor
Submit packages to cran
- http://f.briatte.org/r/submitting-packages-to-cran
- Submitting your first package to CRAN, my experience
- Preparing Your Package for for Submission by John Muschelli
- https://builder.r-hub.io/. See here.
- The most annoying warning for CRAN submission
C/Fortran
Rmath.h
For example pnorm5() was used by survS.h by survival package (old version) .
Packages includes Fortran
On mac, gfortran (6.1) can be downloaded from CRAN. It will be installed onto /usr/local/gfortran. Note that the binary will not be present in PATH. So we need to run the following command to make gfortran avaiialbe.
sudo ln -s /usr/local/gfortran/bin/gfortran /usr/local/bin/gfortran
Misc
Build R package faster using multicore
http://www.rexamine.com/2015/07/speeding-up-r-package-installation-process/
The idea is edit the /lib64/R/etc/Renviron file (where /lib64/R/etc/ is the result to a call to the R.home() function in R) and set:
MAKE='make -j 8' # submit 8 jobs at once
Then build R package as regular, for example,
$ time R CMD INSTALL ~/R/stringi --preclean --configure-args='--disable-pkg-config'
suppressPackageStartupMessages() and .onAttach()
It is Time for CRAN to Ban Package Ads
suppressPackageStartupMessages(library("dplyr"))