Main Page: Difference between revisions
| Line 249: | Line 249: | ||
=== Quirks === | === Quirks === | ||
* Do not place the project in a very deep path. | * Do not place the project in a very deep path. Or you may get an error | ||
<pre> | |||
reads "Error in 'exportToR' function" and then reads "Data was not successfully exported to R. Plug-in script is now aborting." | |||
</pre> | |||
* Do not include special characters (single/double quote, percent sign, etc), in the project name, output name, column header in the experiment descriptor worksheet. The special characters include * ? < > | = + ~ @ # $ % ^ & |. It is defined in <PublicFunctionProcedure.bas/CheckSpecialCharactersReturnBoolean> | * Do not include special characters (single/double quote, percent sign, etc), in the project name, output name, column header in the experiment descriptor worksheet. The special characters include * ? < > | = + ~ @ # $ % ^ & |. It is defined in <PublicFunctionProcedure.bas/CheckSpecialCharactersReturnBoolean> | ||
* Do not sort the experiment descriptor worksheet. | * Do not sort the experiment descriptor worksheet. | ||
Revision as of 13:08, 5 December 2014
Wiki for BRB-ArrayTools .
Features
Import from multiple data types
Expression data, Illumina methylation data, Copy number data (CGH-Tools), RNA-Seq count data processed through Galaxy web tool.
Sophisticated statistical analysis tools
Class comparison for differential expression, class prediction, graphical 2d and 3D interactive plots, gene set analysis, and more.
Comprehensive biological annotations
Gene ontology, pathways, protein domain, broad msigdb, lymphoid signatures, experimentally verified transcription factor targets, computationally predicted microRNA targets.
Screenshots
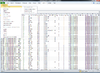 BRB-ArrayTools graphical user interface
BRB-ArrayTools graphical user interface
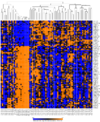 Heatmap and dendrogram generated from the Pomeroy sample dataset.
Heatmap and dendrogram generated from the Pomeroy sample dataset.
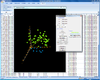 Interactive MDS plot of samples from running the multidimensional scaling analysis on the Pomeroy dataset.
Interactive MDS plot of samples from running the multidimensional scaling analysis on the Pomeroy dataset.
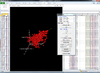 Interactive 3D scatterplot of genes on the Pomeroy dataset.
Interactive 3D scatterplot of genes on the Pomeroy dataset.
X-axis is from array 'Brain_MD_1', y-axis is 'Brain_MD_2' and z-axis is 'Brian_MD_3'.
The right click menu gives an option to highlight up/down-regulated genes, export gene list, copy plot to clipboard, highlight genes in gene set, link genes among plots and change properties of the plot like title, point size, color of points, fold change threshold for up/down regulated genes.
 Interactive volcano plot from the output of running a class comparison tool.
Interactive volcano plot from the output of running a class comparison tool.
When you move mouse over a gene (point), the gene unique ID and/or symbol will be popped up.
 Interactive zoomable heatmap.
Interactive zoomable heatmap.
The program is multi-platform. The screenshot was taken from running in Ubuntu OS. It will be incorporated into BRB-AT in Version 4.4.0.
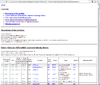 HTML output of running the class comparison analysis.
HTML output of running the class comparison analysis.
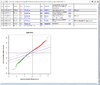 HTML output containing SAM plot from running the significance of microarray analysis.
HTML output containing SAM plot from running the significance of microarray analysis.
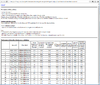 HTML output of running the class prediction analysis.
HTML output of running the class prediction analysis.
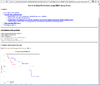 HTML output of running the survival risk analysis.
HTML output of running the survival risk analysis.
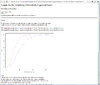 Result of sample size analysis.
Result of sample size analysis.
FAQs
General
How to install BRB-ArrayTools if you have 64-bit MS-Office?
There is no difference in terms of the installation.
After installation, I did not find the BRB-ArrayTools in Windows > Start > All Programs.
Check EXCEL. ArrayTools and CGHTools are under the Excel menu of Addon.
After open Excel, what options I need to do in Excel before using BRB-ArrayTools
Excel -> Home -> Options -> Trust Center -> Trust Center Settings -> Macro Settings -> Check 'Trust access to the VBA project object model' -> OK. No need to restart Excel is required.
Can I upgrade R or install multiple versions of R?
Better not for upgrading. Installing multiple versions of R is OK provided you know some details described below.
Each version of BRB-ArrayTools has been tested with a certain version of R. So there maybe a compatibility problem with certain functions used in the code if you decide to upgrade R.
- BRB-ArrayTools (before v4.3.0) requires StatconnDCOM which means the following conditions have to be satisfied:
- R needs to registered in the Windows's registry (it should be done if you accept all default options when R was installed).
- The R package 'rscproxy' has to be installed under the library folder the registered R. It cannot be installed under user's Document's folder as other R's packages.
- BRB-ArrayTools (from v4.3.0) requires Rserve package. That means
- Rserve has to be installed. It does not have to be installed under R\library folder.
- Rserve.exe from R\librar\Rserve\inst\i386 and x64 subfolder has to be copied to R\bin\i386 and R\bin\x64 folder.
If you need to use the latest version of R for your own analysis, you can still make it. First, install the latest version of R as usual. Then install again the full-version of BRB-ArrayTools. This will possibly install another version of R and register it in the Windows's registry for BRB-ArrayTools to use. Now you can enjoy both versions of R as you want. The idea is when you install R, it will by default register R, but this behavior can break the setting for R to be used by BRB-ArrayTools. Once you install BRB-ArrayTools, it will install an R it needs and not erase any other versions of R you already have.
How to upgrade Bioconductor
For example, my bioc is 2.12 which was first installed when I use R 3.0.1. But bioc 2.13 is the current version when R 3.0.2 was used. When I need to install a new package from bioc, the new package may requires a new version of bioc package.
source("http://bioconductor.org/biocLite.R")
biocLite("BiocUpgrade")
This command will upgrade all currently installed bioc related packages. But it seems it will install lots of other bioc packages I don't need.
My institute is using a proxy server. So how do I do with BRB-ArrayTools
See the message on BRB-ArrayTools message board. Essentially we need to create a Windows environment variable http_proxy with a value like
http://myproxy:myport OR http_proxy=http://username:[email protected] OR http_proxy=http://username:[email protected]:81
RExcel-statconnDCOM gave an error
Please upgrade BRB-ArrayTools to v4.3.x where Rserve has replaced statconnDCOM.
Rserve
Since version 4.3.0, BRB-ArrayTools started to use Rserve as a media for the communication between R and Excel. When Rserve is required, an R window will be pop up. This R window has a blue icon on the Windows' taskbar. If you accidentally close it, it will be automatically popped up when it is needed.
See my Rserve wiki page.
Importing
GEO (Gene Expression Omnibus)
- If there is a GDS number, use GEO importer to import data (eg GDS1348). Expression data, experiment description and gene identifiers will be created automatically. BRB-AT downloads GDSxxx.soft.gz file from this ftp/retired dataset and GPL files from ftp. The <GDSxxxx.soft> file can be used to create <Experiment Descriptor file.txt> and <GDS1348.txt> files for BRB-AT. Note that dataset with GDS number contains experiment info. Individual Experiment (GSMxxxx) does not have experiment info. We can browse all GDS datasets from this ftp link.
- If there is no GDS number,
- If there is raw data, we can try to import them with data import wizard (eg GSE18170). We can use SOURCE to annotation data using 'Symbol' as the lookup key. Select 'Mouse' as the organism.
- If there is no raw data, we can use Serial matrix file with general importer (eg GSE33403). Download GPL10481_family.soft.gz file and Extract it. Open the file in EXCEL. Copy the cells on rows from126 to 45346 and columns from A to I. Paste them to a new Workbook and save it in a tab-delimited format file called <SeriesMatrix.txt>. Download GSE33403_series_matrix.txt.gz file and extract it. Open it in EXCEL. Copy the data on rows from 75 to 45295 and columns from A to AN. Paste them to a new Workbook and save it in a tab-delimited format file called <GeneID.txt>.
Multiple chips (hgu133a and hgu133b)
GSE4922 contains two different array platforms (hgu133a and hgu133b). Starting from ArrayTools v4.1 our software does not support the importing of multi-chips any more. Therefore you cannot directly import all the data into BRB-ArrayTools to create one project with two different chips. However, you could import the data into ArrayTools as two different projects and output the normalized data for each project. Then you can manually combine the two output data files and re-import into BRB-ArrayTools as one project. Here is what you can do,
- Use Data Import Wizard to import the CEL files for hgu133a and hgu133b chips separately to create two projects.
- Open each project and then use the "Export 1-color data to R" plug-in (click on "ArrayTools -> Plugins -> Export 1 color data to R") to output your normalized data file along with the GeneId file.
- Under Excel, combine the normalized output files ("NORMALIZEDLOGINTENSITY.txt") from two projects. Add sample names in the first row and Probe Set Ids in the first column. Save this combined data file. Then combine the two "GENEID.txt" files manually to create one Gene ID file.
- Open Excel, Click on "ArrayTools -> Import data -> General format importer" to import the combined data file. Select your data as "Single-channel", "Affymetrix probeset-summary data". For the chip type, because your data contain probe sets from both hgu133a and hgu133b, almost all of which were included in the hgu133plus2 chip, you can just pick the hgu133plus2 as your chip type. Alternatively, if you do not wish to do this, you can check "I would like to use my own gene identifiers file rather than the one from bioconductor" and use the combined Gene ID file (done in step 3) for annotation.
- At the filter and normalization step, it is VERY IMPORTANT that you need to uncheck all the spot filter and normalization options, because your "raw" data file comes from already normalized data in two different projects. You do not want to re-run normalization. You can keep the options in "Gene filter" tab.
- At the end you can run annotation as your choice.
PrimeView importing failed on v4.3.0 beta1 (R 2.14.x)
- Go to http://www.bioconductor.org/packages/2.11/data/annotation/html/primeviewcdf.html and click on the link for the windows binary package "primeviewcdf_2.11.0.zip" and download the zipped package.
- Unzip the downloaded package and put the entire package folder "primeviewcdf" under your R2.14.2 library folder (default C:\Program Files\R\R-2.14.2\library).
However, you cannot run Affymetrix annotation using bioconductor annotation packages at the end of importing, because the annotation package for primeview array is not available at bioconductor. Another commonly use annotation database, SOURCE, has also been down these days. Instead you need to import your own annotation into BRB-ArrayTools. Here is what you can do,
- Download the primeview annotation file in CSV format from Affymetrix website (https://www.affymetrix.com/user/login.jsp?toURL=/support/file_download.affx?onloadforward=/analysis/downloads/na32/ivt/PrimeView.na32.annot.csv.zip).
- Open the CSV format file in Excel and then save it as a tab-delimited .txt file.
- Import your data in CEL file format into BRB-ArrayTools through Data Import Wizard. During importing, you need to check the option "Import your own identifiers file to annotate your data" at the "Probe Set Id options" page, and then Click on "OK". At the next dialog form you need to select the radio button "The identifiers are stored in a separate file" and browse for the tab-delimited file you just saved. The head line # for this file is 25. Then you need to select the corresponding gene identifiers column headers. You need to check the box "Annotate the project with these gene ids, instead of using the data from SOURCE database" before proceeding to the next page.
Affymetrix miRNA 3.0
For Affymetrix miRNA 2.0 array (not for 3.0 array since there is no cdf package), you can import the data in CEL file format into BRB-ArrayTools through the CEL file importer (from Data Import Wizard). For miRNA 2.0 and 3.0 arrays, an alternative way is to use the Affymetrix expression console software to pre-process/normalize the CEL file data and obtain a tab-delimited probe-set summary data file to be imported into BRB-ArrayTools using the General format importer. The Affymetrix expression console software can be downloaded at the Affymetrix website (http://www.affymetrix.com/estore/browse/level_seven_software_products_only.jsp?navMode=34000&productId=131414&navAction=jump&aId=productsNav#1_1). Unlike what we do for the regular gene expression chips, BRB-ArrayTools currently does not support the generation of annotation information based on the microRNA ID (so gene set analysis cannot be run).
For now, the CSV format annotation file provided by Affymetrix can be opened in Excel and then saved as a tab-delimited .txt file, which can be used as a "Gene Identifiers" file for BRB-ArrayTools. When you import your data file into BRB, you can choose to "use your own gene ID file for annotation" and select the "Transcript Id (Array Design)" as the microRNA Id column header. Although we do not directly provide any annotation tools for miRNA data, if you run some particular analyses such as class comparison, there will be a hyperlink associated with the microRNA Id in your html output result file. By clicking on this link, you can be directed to the miRBase website with the annotation information associated with this particular microRNA Id.
Agilent miRNA
The arrays in the GSE41874 series were scanned by an Agilent scanner and data were extracted by the feature extraction software, therefore the raw data files in this series are in the exactly same format as the Agilent dual-channel .txt files. You can use Data Import Wizard and choose "Agilent dual-channel data" to import the data into BRB-ArrayTools. However, you need to select the following options:
- At the GeneID dialog form page, select the "GeneName" as the miroRNA ID, and select "None" as the Gene Symbol
- At the spot filtering step, use dye swap for all the arrays, as Cy5 was used as the reference for all arrays
- Also, please bear in mind that only some analysis tools can be used for this set of data, because annotation cannot be conducted on microRNA array data in BRB-ArrayTools. For instance, you can run class comparison and clustering with this data set, but cannot perform gene set enrichment analysis.
Although annotation is not provided, at the end of class comparison, the significant microRNAs can be hyperlinked to the mirBase if you want to see detailed annotation of any particular microRNAs.
Gene ST 1.1 (or other than 1.0), Exon ST, or Affymetrix .cel files without cdf packages from Bioconductor
You will need to install Affymetrix Expression Console. Here is the link of Affymetrix Expression Console download website. http://www.affymetrix.com/browse/level_seven_software_products_only.jsp?productId=131414&categoryId=35623#1_1
Here is the link where you could download Affymetrix Expression Console (64bit).
Take the mouse Gene 1.1 ST-array as an example on how to use the Expreesion console to convert .cel files into .txt files and then import .txt files into ArrayTools.
- Before downloading the Affymetrix expression console, you need to register at Affymetrix website. After downloading and installing the software, open Affymetrix expression console, name the profile and set a library path by clicking "Edit -> Set library path";
- After you download and install the software, please open it and then download both the library file and annotation file for mogene 1.1 ST-array in the software(File->Download Library Files and File->Download Annotation Files).
- After you download both files, you could click on "File" -> "New study" to open the Affymetrix study window. Click on "Add intensity files" to browse for the CEL files of your interest. Click on "Run analysis".
- When the analysis is done, click on "Export" -> "Export Probe Set Results(pivot table) to TXT" to export a tab-delimited file, which is ready to be imported into BRB-ArrayTools.
- Go to the library path folder and open a file called "xxxx.annot.csv" in Excel. Save this file in tab-delimited .txt format as an annotation file for later use.
- You could start to use BRB-ArrayTools General importer to read the summarization .txt file into ArrayTools. By default, the output expression values from Affymetrix Expression Console is already log 2 transformed. Please turn off all spot filters and normalization during importing in ArrayTools because RMA is already done by Affymetrix Expression Console.
- At the Gene ID dialog form, select "Gene Ids are stored in a separate file" and browse for the annotation file you created in step 5. Check the option "Annotate the project with these gene ids, instead of using the data from SOURCE database".
Annotation
SOURCE at stanford is down
During the period of time when SOURCE website is down, our users will not be able to import annotations from SOURCE database. The bioconductor annotation packages will still work for users having Affymetrix or Illumina array data. For non-Affymetrix and non-Illumina chip users who are creating new projects, here are a couple of alternative solutions,
- If the user has an existing annotated project with an identical chip type to the project he/she is creating, he/she can import the annotations from the existing project. This can be done by clicking on 'ArrayTools -> Utilities -> Annotate the data -> Import annotations from an existing project';
- If the user has an annotation file available, during the process of importing, the user can choose to import annotations from this file. At the step of selecting Gene Identifiers, the user can browse for the Gene Identifiers file and check the option 'Annotate the project with these gene ids, instead of using the data from SOURCE database';
- If the user does not have an annotation file, for most commonly used commercial chips, the annotation files can be downloaded from GEO database at NCBI. For instance, the annotation file for the Agilent-014850 Whole Human Genome Microarray chip can be downloaded at http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GPL4133 by clicking on the 'Download full table' button. This downloaded annotation file can then be imported into BRB-ArrayTools following step 2).
Gene ST array annotation using aroma.affymetrix
The aroma.affymetrix can handle all single-channel Affymetrix chip types (they have a few multi-channel ones not supported), so my guess is that it is already supported. What's required for nearly all aroma analysis, is to have a CDF annotation file that defines the units and unit groups, e.g. probe sets of transcripts and exons, SNPs and so on. When Affymetrix is not providing a CDF, the challenge is to either find a custom CDF (e.g. BrainArray and GeneAnnot) or to create a one from the other types of annotation data they or Bioconductor provide, e.g. http://aroma-project.org/howtos.
Annotation with Bioconductor package
The tool search gene symbol, accession number, UGCluster and Entrez ID columns. See <AnnotationGenes.bas> for the source code.
An Example of a Gene Annotation
| CloneID | 50188 |
| LLID (=Entrez ID or GeneID) | 6047 |
| ProbeID | AB000468_at |
| Symbol | RNF4 |
| UGCluster | Hs.66394 |
| RefSeq (=accession number) | NM_000844 |
| GenBank Accession | M97935 |
| GO | nucleus|TAS|GO:0005634//cellular component|nucleus|IEA|GO:0005634 |
Analysis
Function analysis using DAVID
See a tutorial from http://nihlibrary.ors.nih.gov/bioinfo/Microarray/Problem1.html
Best parameter or analysis to choose
Ask your local help desk.
For affy cel files, GCRMA has a memory problem and MAS5 is way too slow
It is a known problem for earlier versions of BRB-ArrayTools. The reason is statconnDCOM can only use 32-bit R. Since BRB-ArrayTools v4.3.0, a new engine 'Rserve' was adopted which can make use 64-bit R if the Windows OS is 64-bit. Therefore the memory issue for using GCRMA should be alleviated. (Update) The 64-bit OS still does not solve the memory problem; see a post and a workaround here from google forum.
BRB-ArrayTools uses justGCRMA() function in 'gcrma' package for GCRMA option and justMAS() function in 'simpleaffy' package for MAS5 option.
GO
- Gene annotation worksheet is used in Utilities -> Create genelist -> GO description.
- Bioconductor's GO package is used when we run gene set (gene ongology) class comparison.
R packages
- GCRMA: install cdf, probe, .db packages
- MAS5: install cdf, .db
Common download issue
Try to download/install manually in R console.
- lumi
source("http://bioconductor.org/biocLite.R")
biocLite("lumi", ask=F)
library(lumi)
- GO.db
source("http://bioconductor.org/biocLite.R")
biocLite("GO.db", ask=F)
Users without Admin Privilege
- log in as the administrator and set up BRB-ArrayTools in Excel. Plus, I had to enter my email address for activation.
- log in as standard user and click the Go button for Manage Excel add-ins. Since BRB-ArrayTools add-in was still not showing up in the dialog box, I had to manually add it by clicking on the Browse button.
Foreign langauge users
Below are some of the recommendations we typical make to foreign language users: 1: Please, make sure that the regional language settings on your machine to the "English". You can do so by going to the Control Panel -> Regional and Language Options and choose English. 2: From the Start Programs Microsoft office Microsoft office tools -> ->"Microsoft Office language settings" and please make sure that the "primary editing language" is "English". If the "Primary editing language" is not "English", please change it to "English" and then re-boot your machine.
If we don't want to change the setting to "English". The trick was to: in Excel Add-Ins, look for the add-ins among folders (in the German version: Durchsuchen) and to folder programs v 64 bit. Then it finds ArrayTools there, go to subfolder Excel and there it finds the Add-in. Otherwise it was invisible.
In the administrator system, allow British English keyboard,
And in Excel Options -> Advanced -> Change decimal to dot . and thousands to comma.
Quirks
- Do not place the project in a very deep path. Or you may get an error
reads "Error in 'exportToR' function" and then reads "Data was not successfully exported to R. Plug-in script is now aborting."
- Do not include special characters (single/double quote, percent sign, etc), in the project name, output name, column header in the experiment descriptor worksheet. The special characters include * ? < > | = + ~ @ # $ % ^ & |. It is defined in <PublicFunctionProcedure.bas/CheckSpecialCharactersReturnBoolean>
- Do not sort the experiment descriptor worksheet.
- R's impute package tends to crash R when the number of genes is small.
- R's pamr package failed when the number of genes is only one. The error message is
Error in rep(1, p) : invalid 'times' argument
It is a bug in pamr.train() -> nsc().
- Write the R file used in plugins in a conventional format.
- Bioconductor package 'affxparser' does not work on Windows XP. The alternative is to use Affymetrix Expression Console to pre-process your ST arrays data and then import the .txt file that are outputted from Affymetrix Expression Console into ArrayTools by using the general format importer.
- Sometimes we need to delete the parameter file (under $ProjectFolder\BinaryData\DataParam) to solve a problem. For example, two projects were opened at the same time, or an analysis was broken during execution.
Run time errors
This is a collection of run-time errors from users' report or testings.
- error 1004: make sure the specified folder exists or file can be accessed.
- error 91: 'block variable not set'. It is rscproxy is not installed correctly. This should not happen again since rcom is no longer used.
- error 76: write permission, administrative privileges.
- error 75: write permission
- error 9: subscription out of range. Variable name used in dialog is changed.
- error 5: invalid procedure call or argument.
- error '-2147319779' or '-2147221500': rscproxy package is not installed. This should not happen again since rcom is no longer used.
Citing
Plugins Developers
- Experiment descriptor is a data frame with numerical or character data type (no factor). So we shall take extra care for cell with NA (numerical data type) or blank (character data type) value.
- Gene identifier is a data frame with factor data type.
Support
Send an email to [email protected]
Please provide enough information to us so we can understand the problem.
- If a bug report file was generated, be sure to send it to us.
- If the question is like 'what method or parameters should be choose to run my analysis', please consult other experienced people near you.
- Since the software depends on a couple factors like Windows operation system, MS-Office, R. Please provide us more detailed information about the software background including BRB-ArrayTools.
- When sending screenshots to us, please provide all error screenshots. If you only provide any random of them, it will create a misleading to us.
Check BRB-ArrayTools message board [1]
MISC
Illumina Genome Studio Manual
GEO
- Illumina. For example, the txt file <GSE13040_nonorm_nobkgd.txt> from GSE13040 (Illumina MouseRef-8 v2.0 expression beadchip) is close but not the one BRB-ArrayTools requires. We can modify the header to satisfy AT's requirement.
Affymetrix SNP arrays and cnt file
To use the ‘Platform special importer’ for Affymetrix SNP arrays, the user should run the Copy Number Analysis Tool (CNAT) software provided freely by Affymetrix, and output the *.CNT files in a batch process, where one *.CNT file is produced for each SNP array that was performed. The *.CNT output files should all be placed in one data folder, to be read by CGHTools. The “ProbeSet”, “Chromosome”, “Position” and “Log2Ratio” data columns will be automatically extracted from each *.CNT file.
See also goldenhelix.com for more about creating CNT files using the Affymetrix CNAT Batch Analysis Tool.
Affymetrix website contains some info about Chromosome Copy Number Analysis Tool (CNAT) software. This site provides links to command line tools to process 10K, 100K, and 500K (no SNP 5 or SNP 6) data from CHP file to CNT file. In fact, the tool download page here even provides sample output for download.
We may obtain raw files of copy number data from GEO website by searching 'affymetrix' and '10k' under 'Platforms' tab as keywords.
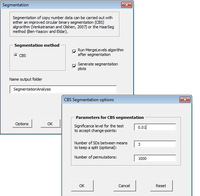
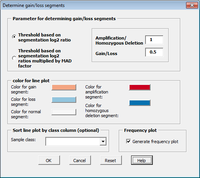
CGHTools
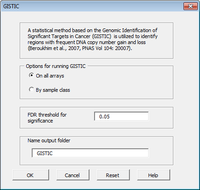
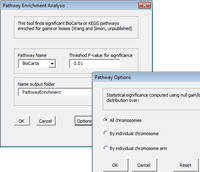
The CGHTools has several tools (segmentation, gain/loss, Gistic, pathway).
Once segmentation has been run, it is OK to jump to gain/loss, Gistic or pathway analysis. These 3 analyses have no mutual dependencies.
The CGHTools manual said when inferred integer copy number is imported at the importing step, the pathway enrichment analysis can be conducted without segmentation being performed.