Venn diagram: Difference between revisions
Appearance
| (One intermediate revision by the same user not shown) | |||
| Line 50: | Line 50: | ||
* [https://cran.r-project.org/web/packages/ComplexUpset/index.html ComplexUpset]: Create Complex UpSet Plots Using 'ggplot2' Components | * [https://cran.r-project.org/web/packages/ComplexUpset/index.html ComplexUpset]: Create Complex UpSet Plots Using 'ggplot2' Components | ||
* [https://jokergoo.github.io/ComplexHeatmap-reference/book/upset-plot.html Chapter 8 UpSet plot] | * [https://jokergoo.github.io/ComplexHeatmap-reference/book/upset-plot.html Chapter 8 UpSet plot] | ||
= VennDiagram* = | |||
<ul> | |||
<li>[https://cran.r-project.org/web/packages/VennDiagram/ VennDiagram]. As good as the "venn" package. Many reverse imports and suggests. | |||
<li>[https://www.datanovia.com/en/blog/venn-diagram-with-r-or-rstudio-a-million-ways/#using-the-venndiagram-r-package Venn Diagram with R or RStudio: A Million Ways]. Question: why the sets orders are changed? | |||
<li>Example from the [https://www.rdocumentation.org/packages/VennDiagram/versions/1.8.2/topics/venn.diagram help] | |||
<syntaxhighlight lang='r'> | |||
grid.newpage() | |||
grid.draw(venn.diagram(list(A = 1:150, B = 121:170), NULL) ) # no colors (BW), circle size proportional to size | |||
list1 <- paste0("A", 1:30) | |||
list2 <- c(paste0("A", 15:35), paste0("B", 1:10)) | |||
list3 <- c(paste0("A", 10:20), paste0("B", 5:15), paste0("C", 1:10)) | |||
grid.newpage() | |||
grid.draw(venn.diagram(list(A = list1, B = list2, C=list3), NULL, | |||
category.names = c("Set 1" , "Set 2 ", "Set 3"), | |||
fill = c("#0073C2FF", "#CD534CFF", "#EFC000FF"), | |||
alpha = .8, | |||
col = "transparent" # no border | |||
) ) # 3 circles same size | |||
</syntaxhighlight> | |||
</li> | |||
<li>[https://bmcbioinformatics.biomedcentral.com/articles/10.1186/1471-2105-12-35 VennDiagram]: a package for the generation of highly-customizable Venn and Euler diagrams in R. scaling is one option. </li> | |||
<li>https://www.r-graph-gallery.com/venn-diagram.html </li> | |||
<li>And a [https://stackoverflow.com/a/11935618 trick] to make scaling works. </li> | |||
</ul> | |||
= venn = | = venn = | ||
| Line 69: | Line 95: | ||
https://cloud.r-project.org/web/packages/ggVennDiagram/index.html, [https://gaospecial.github.io/ggVennDiagram/ github]. | https://cloud.r-project.org/web/packages/ggVennDiagram/index.html, [https://gaospecial.github.io/ggVennDiagram/ github]. | ||
<pre> | |||
# Install if needed | |||
install.packages("ggVennDiagram") | |||
library(ggVennDiagram) | |||
library(ggplot2) | |||
# Define your sets | |||
genes <- list( | |||
GroupA = c("gene1", "gene2", "gene3", "gene4", "gene5"), | |||
GroupB = c("gene3", "gene4", "gene6", "gene7"), | |||
< | GroupC = c("gene4", "gene5", "gene7", "gene8", "gene9") | ||
) | |||
# Basic plot | |||
ggVennDiagram(genes) + | |||
scale_fill_gradient(low = "#F4FAFE", high = "#08519C") + | |||
theme(legend.position = "none") | |||
# Save publication-ready figure | |||
ggsave("venn_diagram.pdf", width = 6, height = 5, dpi = 300) | |||
ggsave("venn_diagram.png", width = 6, height = 5, dpi = 300) | |||
</pre> | </pre> | ||
= ggvenn = | = ggvenn = | ||
| Line 127: | Line 155: | ||
= eulerr = | = eulerr = | ||
[https://cran.r-project.org/web/packages/eulerr/index.html eulerr] package | * [https://cran.r-project.org/web/packages/eulerr/index.html eulerr] package | ||
* The circle sizes are proportional to the list sizes. However, the diagram can look very weird sometimes. | |||
* For some reason, if I use png() to save the plot, it works in the interactive mode, but not when I try to render a Rmd file. | |||
Latest revision as of 11:33, 26 February 2026
Misc
- 2014 Paper eulerAPE: Drawing Area-Proportional 3-Venn Diagrams Using Ellipses which includes an overview of several tools. The paper is linked from the website https://upset.app.
- Visualization biological data using proportional venn diagrams
- VENN DIAGRAM WITH R OR RSTUDIO: A MILLION WAYS
- http://manuals.bioinformatics.ucr.edu/home/R_BioCondManual#TOC-Venn-Diagrams and the R code or the Bioc package systemPipeR
# systemPipeR package method library(systemPipeR) setlist <- list(A=sample(letters, 18), B=sample(letters, 16), C=sample(letters, 20), D=sample(letters, 22), E=sample(letters, 18)) OLlist <- overLapper(setlist[1:3], type="vennsets") vennPlot(list(OLlist)) # R script source method source("http://faculty.ucr.edu/~tgirke/Documents/R_BioCond/My_R_Scripts/overLapper.R") setlist <- list(A=sample(letters, 18), B=sample(letters, 16), C=sample(letters, 20), D=sample(letters, 22), E=sample(letters, 18)) # or (obtained by dput(setlist)) setlist <- structure(list(A = c("o", "h", "u", "p", "i", "s", "a", "w", "b", "z", "n", "c", "k", "j", "y", "m", "t", "q"), B = c("h", "r", "x", "y", "b", "t", "d", "o", "m", "q", "g", "v", "c", "u", "f", "z"), C = c("b", "e", "t", "u", "s", "j", "o", "k", "d", "l", "g", "i", "w", "n", "p", "a", "y", "x", "m", "z"), D = c("f", "g", "b", "k", "j", "m", "e", "q", "i", "d", "o", "l", "c", "t", "x", "r", "s", "u", "w", "a", "z", "n"), E = c("u", "w", "o", "k", "n", "h", "p", "z", "l", "m", "r", "d", "q", "s", "x", "b", "v", "t"), F = c("o", "j", "r", "c", "l", "l", "u", "b", "f", "d", "u", "m", "y", "t", "y", "s", "a", "g", "t", "m", "x", "m" )), .Names = c("A", "B", "C", "D", "E", "F")) OLlist <- overLapper(setlist[1:3], type="vennsets") counts <- list(sapply(OLlist$Venn_List, length)) vennPlot(counts=counts)
Online tools
- https://molbiotools.com/listcompare.php
- https://www.interactivenn.net/
- https://www.deepvenn.com/.
- Proportional Circles
- Labels (eg "Set A") can be moved outside manually if we click "Draw as SVG"
- It has a 4 built-in examples (near the bottom)
UpSet plot
- https://upset.app/
- UpSetR
- ComplexUpset: Create Complex UpSet Plots Using 'ggplot2' Components
- Chapter 8 UpSet plot
VennDiagram*
- VennDiagram. As good as the "venn" package. Many reverse imports and suggests.
- Venn Diagram with R or RStudio: A Million Ways. Question: why the sets orders are changed?
- Example from the help
grid.newpage() grid.draw(venn.diagram(list(A = 1:150, B = 121:170), NULL) ) # no colors (BW), circle size proportional to size list1 <- paste0("A", 1:30) list2 <- c(paste0("A", 15:35), paste0("B", 1:10)) list3 <- c(paste0("A", 10:20), paste0("B", 5:15), paste0("C", 1:10)) grid.newpage() grid.draw(venn.diagram(list(A = list1, B = list2, C=list3), NULL, category.names = c("Set 1" , "Set 2 ", "Set 3"), fill = c("#0073C2FF", "#CD534CFF", "#EFC000FF"), alpha = .8, col = "transparent" # no border ) ) # 3 circles same size - VennDiagram: a package for the generation of highly-customizable Venn and Euler diagrams in R. scaling is one option.
- https://www.r-graph-gallery.com/venn-diagram.html
- And a trick to make scaling works.
venn
- venn package (best so far)
- polygon() function call.
- Introduction to the venn Package in R (6 Examples) | How to Draw Up to 7 Sets
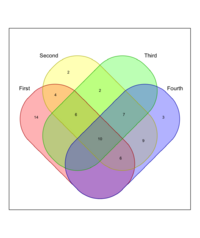
library(venn)
set.seed(12345)
x <- list(First = 1:40, Second = 15:60, Third = sample(25:50, 25),
Fourth=sample(15:65, 35))
venn(x, ilabels = "counts", zcolor = "style", ilcs = 1.2, sncs = 1)
In this example, ilcs and sncs will control the font size of the intersection labels and set names, respectively.
ggVennDiagram
https://cloud.r-project.org/web/packages/ggVennDiagram/index.html, github.
# Install if needed
install.packages("ggVennDiagram")
library(ggVennDiagram)
library(ggplot2)
# Define your sets
genes <- list(
GroupA = c("gene1", "gene2", "gene3", "gene4", "gene5"),
GroupB = c("gene3", "gene4", "gene6", "gene7"),
GroupC = c("gene4", "gene5", "gene7", "gene8", "gene9")
)
# Basic plot
ggVennDiagram(genes) +
scale_fill_gradient(low = "#F4FAFE", high = "#08519C") +
theme(legend.position = "none")
# Save publication-ready figure
ggsave("venn_diagram.pdf", width = 6, height = 5, dpi = 300)
ggsave("venn_diagram.png", width = 6, height = 5, dpi = 300)
ggvenn
- ggvenn package
- RNA-seq Analysis in R
library(ggvenn)
ggvenn(list(A = 1:150, B = 121:170, C=141:200),
fill_color = c("#0073C2FF", "#CD534CFF", "#EFC000FF")) # looks normal
Cons: The generated plot looks good. But the locations of set labels are weird and not adjustable.
gplots
black and white?
gplots::venn(list(A = 1:150, B = 121:170, C=141:200))
Vennerable
Vennerable package https://github.com/js229/Vennerable
- Make Venn Diagram in R with Correctly Weighted Areas
- Venn Diagrams with R
if (!requireNamespace("BiocManager", quietly = TRUE)) install.packages("BiocManager") BiocManager::install("graph") BiocManager::install("RBGL") # fail on Rstudio cloud install.packages("Vennerable", repos="http://R-Forge.R-project.org")
limma
- https://rdrr.io/bioc/limma/man/venn.html
- User guide. Only black and white.
eulerr
- eulerr package
- The circle sizes are proportional to the list sizes. However, the diagram can look very weird sometimes.
- For some reason, if I use png() to save the plot, it works in the interactive mode, but not when I try to render a Rmd file.