Search results
Jump to navigation
Jump to search

File:Cvglmnetplot.png set.seed(1) # [1] 39(1,000 × 400 (34 KB)) - 17:56, 11 October 2022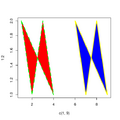
File:Polygon.png plot(c(1, 9), 1:2, type = "n") polygon(1:9, c(2,1,2,1,NA,2,1,2,1),(577 × 577 (30 KB)) - 17:31, 23 April 2024
File:Logscale.png > plot(c(1, 10, 100), c(1, 10, 100), log="y") > log10(c(1, 10, 100))(480 × 480 (4 KB)) - 13:58, 1 May 2020
File:Jitterbox.png assy <- LETTERS[1:nc] set.seed(1)(1,020 × 906 (50 KB)) - 22:53, 8 February 2024
File:Reorder.dendrogram.png par(mfrow=c(3, 1)) plot(dd, main = "random dendrogram 'dd'") # not the same as reorder(dd, 1:20)(814 × 600 (10 KB)) - 12:15, 8 January 2023
File:ViridisDefault.png 1:n, 1, as.matrix(1:n),(600 × 200 (2 KB)) - 10:31, 30 August 2022
File:Twopalette.svg par(mfrow=c(1,2), mar = c(5,2,2,1)) plot(1:8, col=brewer_pal(palette = "Blues")(8), pch=20, cex=4)(468 × 468 (43 KB)) - 16:09, 12 April 2019
File:Methybetavsm.png mydf = data.frame(beta=seq(0,1,length=100), m=log2(seq(0,1,len=100)/(1-seq(0,1,len=100))))(480 × 480 (3 KB)) - 10:21, 30 July 2015
File:Scatterhist.svg zones=matrix(c(2,0,1,3), ncol=2, byrow=TRUE) layout(zones, widths=c(4/5,1/5), heights=c(1/5,4/5))(563 × 563 (70 KB)) - 10:22, 16 February 2024
File:Rpalette.png # [1] "black" "#DF536B" "#61D04F" "#2297E6" "#28E2E5" "#CD0BBC" "#F5C710" pal_matrix <- matrix(seq_along(pal), nr=1)(534 × 615 (28 KB)) - 11:08, 10 May 2023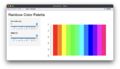
File:Rainbow default.png sliderInput("s_value", "Saturation (s):", min = 0, max = 1, value = 1, step = 0.01), sliderInput("v_value", "Value (v):", min = 0, max = 1, value = 1, step = 0.01)(2,044 × 1,182 (426 KB)) - 08:52, 12 March 2024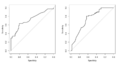
File:Roc asah.png par(mfrow=c(1,2)) par(mfrow=c(1,1))(1,024 × 535 (38 KB)) - 15:59, 22 August 2023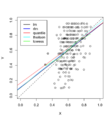
File:DataOutliers.png plot(puree[,1], puree[, 2], xlim=c(0,1), ylim=c(0,1), xlab="X", ylab="Y") abline(0,1, lty=2)(1,020 × 1,096 (172 KB)) - 15:27, 7 March 2024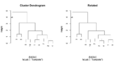
File:Dend12.png # [1] 8 3 6 5 10 1 9 7 2 4 # Rotate the branches (1,9) & (7,2,4)(876 × 448 (11 KB)) - 13:17, 13 August 2023
File:Rotateheatmap.png rownames(mat) = paste0("R", 1:8) colnames(mat) = paste0("C", 1:10)(1,314 × 448 (52 KB)) - 14:44, 13 August 2023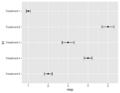
File:Geomerrorbarh.png trt = factor(c("Treatment 1", "Treatment 2", "Treatment 3", "Treatment 4", "Treatment 5")), # treatment resp = c(1, 5, 3, 4, 2), # response(516 × 400 (17 KB)) - 20:38, 15 October 2023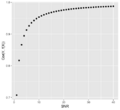
File:SnrVScor.png tibble(snr = seq(1,40)) %>% mutate(cor=sqrt(snr)/sqrt(snr+1)) %>%(483 × 443 (20 KB)) - 14:15, 18 March 2022
File:ComplexHeatmap2.png set.seed(1) colnames(mat) <- 1:ns(800 × 600 (43 KB)) - 21:37, 7 January 2023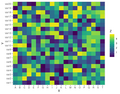
File:ScaleFillViridisDiscrete.png x <- LETTERS[1:20] y <- paste0("var", seq(1,20))(600 × 471 (22 KB)) - 10:08, 30 August 2022
File:GseaTable2.png fgseaRes[order(pval, decreasing = T),][1:10, c('NES', 'pval')] # 1: -0.4050950 1.0000000(800 × 600 (81 KB)) - 12:18, 12 January 2023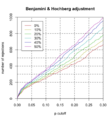
File:Filtered p.png # Each curve represents theta (filtering threshold). For example, theta=.1 means 10% of genes are filtered out before we do multiple testing (or BH ad #* if theta=0, 251 hypotheses are rejected at FDR=.1(1,020 × 1,096 (171 KB)) - 21:35, 11 March 2024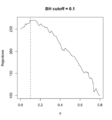
File:Filtered R mean.png # 10% <---- so theta=0.1 is the optimal; only 10% genes are removed # [1] 270(1,020 × 1,096 (68 KB)) - 20:17, 12 March 2024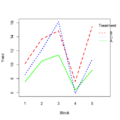
File:RbdBlock.png block <- as.factor(rep(1:5, each=6)) random_effect <- rnorm(30, mean = 0, sd = 1)(500 × 500 (34 KB)) - 21:51, 14 January 2023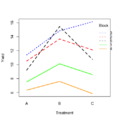
File:RbdTreat.png block <- as.factor(rep(1:5, each=6)) random_effect <- rnorm(30, mean = 0, sd = 1)(500 × 500 (32 KB)) - 21:50, 14 January 2023
File:Geom boxplot.png ggplot(data.frame(x), aes(1, x)) + geom_boxplot()(436 × 536 (14 KB)) - 16:02, 25 November 2019
File:Geom bar2.png y = c(.02, .11, -.01, -.03, -.03, .02, .1, -.01, -.02, 0.06)) set.seed(1)(2,400 × 1,800 (71 KB)) - 14:41, 15 December 2022
File:Geom bar1.png y = c(.02, .11, -.01, -.03, -.03, .02, .1, -.01, -.02, 0.06)) set.seed(1)(2,400 × 1,800 (85 KB)) - 14:39, 15 December 2022
File:KMannotation.png # [,1] [,2] [,3] [,4] [,5] [,6] [,7] [,8] [,9] [,10] [,11] # status 1 1 0 1 1 0 1 1 0 1 0(732 × 546 (58 KB)) - 22:12, 13 October 2020
File:IGV fly Anders2013 2.png Note that a position may contain more than 1 transcript.(1,033 × 854 (102 KB)) - 14:50, 21 May 2015
File:Geom bar4.png color = "grey", linetype = "dashed", size = 1) +(2,400 × 1,800 (51 KB)) - 15:14, 15 December 2022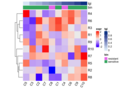
File:ComplexHeatmap1.png rownames(mat) = paste0("R", 1:ng) colnames(mat) = paste0("C", 1:n)(600 × 450 (29 KB)) - 15:02, 23 August 2022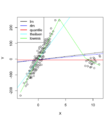
File:DataOutliers2.png plot(puree[,1], puree[, 2], xlab="X", ylab="Y") col = c("black", "blue", "red", "cyan", "green"), lty=1, lwd=2)(1,020 × 1,096 (133 KB)) - 16:49, 8 March 2024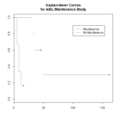
File:KMcurve.png # https://www.rdocumentation.org/packages/survival/versions/3.1-12/topics/print.survfit 1 Maintained 45(672 × 671 (6 KB)) - 16:15, 30 April 2020
File:Svg4.svg plot(1:10, main="width=4, height=4")(360 × 360 (33 KB)) - 16:58, 17 March 2023
File:Palettebarplot.png barplot(rep(1, length(pal)), col = pal, space = 0,(534 × 615 (9 KB)) - 11:15, 10 May 2023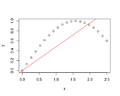
File:R2.png # [1] 0.8554949(500 × 400 (15 KB)) - 13:46, 21 May 2023
File:Venn4.png x <- list(First = 1:40, Second = 15:60, Third = sample(25:50, 25),(982 × 1,168 (103 KB)) - 14:49, 23 April 2024
File:Rerrorbar.svg geom_jitter(position = position_jitter(0.2, seed=1), color = "darkgray") + geom_jitter(position = position_jitter(0.2, seed=1), color = "darkgray") +(609 × 328 (75 KB)) - 19:37, 24 March 2022
File:Rerrorbars.png geom_jitter(position = position_jitter(0.2, seed=1), color = "darkgray") + geom_jitter(position = position_jitter(0.2, seed=1), color = "darkgray") +(593 × 418 (10 KB)) - 19:39, 24 March 2022
File:LogisticFail.svg > set.seed(1234); n <- 16; mu=3; x <- c(rnorm(n), rnorm(n, mu)); y <- rep(0:1, each=n)(475 × 468 (25 KB)) - 10:08, 26 May 2022
File:Fgsea3plots.png par(mfrow=c(1,3))(1,200 × 400 (68 KB)) - 12:37, 6 January 2023
File:Filtered R.png # [1] 380 main="BH cutoff = 0.1")(1,020 × 1,096 (77 KB)) - 21:41, 11 March 2024
File:Foldchangefilter.png LFC <- log2(1.5) plot(x, y = 1:28, xlab="log2 intensity", ylab="samples")(460 × 460 (22 KB)) - 11:03, 12 October 2022
File:Geomcolviridis.png pop_df2 <- tibble(class=rbinom(nrow(pop_df), 1, .5), pop_df)(821 × 400 (26 KB)) - 11:50, 30 August 2022
File:Doubledip.png set.seed(1) i1 <- i[1] ; i2 <- i[2](800 × 450 (57 KB)) - 14:19, 22 August 2022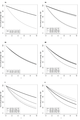
File:Progpreg.png x1 <- image_read(u[1])(800 × 1,200 (193 KB)) - 10:48, 24 September 2020
File:GseaTable.png # 1: 5992282_ECM_proteoglycans 1.984081 # 2: 5992219_Regulation_of_cholesterol_biosynthesis_by_SREBP_SREBF_ 1.955961(800 × 600 (81 KB)) - 12:01, 12 January 2023
File:Ruspini.png mgp= c(1.6,.8,0), mar= .1+c(4,2,2,2))(1,000 × 550 (81 KB)) - 10:41, 10 August 2022