Anders2013: Difference between revisions
| Line 334: | Line 334: | ||
=== [http://ccb.jhu.edu/software/tophat/manual.shtml Tophat options] === | === [http://ccb.jhu.edu/software/tophat/manual.shtml Tophat options] === | ||
* --coverage-search/--no-coverage-search: According to [https://www.biostars.org/p/49224/ this], it is a step to define possible junctions between exons. If you only want the expression profile of the annotated genes, you can skip this step for speed. | * --coverage-search/--no-coverage-search: According to [https://www.biostars.org/p/49224/ this], it is a step to define possible junctions between exons. If you only want the expression profile of the annotated genes, you can skip this step for speed. The identification of new regions with the coverage is relevant only if you want to detect new splice sites in alternate transcripts or even new genes. | ||
The identification of new regions with the coverage is relevant only if you want to detect new splice sites in alternate transcripts or even new genes. | |||
* -r/--mate-inner-dist <int>: This is the expected (mean) inner distance between mate pairs. For, example, for paired end runs with fragments selected at 300bp, where each end is 50bp, you should set -r to be 200. The default is 50bp. | * -r/--mate-inner-dist <int>: This is the expected (mean) inner distance between mate pairs. For, example, for paired end runs with fragments selected at 300bp, where each end is 50bp, you should set -r to be 200. The default is 50bp. | ||
* -g/--max-multihits <int>: Instructs TopHat to allow up to this many alignments to the reference for a given read, and choose the alignments based on their alignment scores if there are more than this number. The default is 20 for read mapping. | * -g/--max-multihits <int>: Instructs TopHat to allow up to this many alignments to the reference for a given read, and choose the alignments based on their alignment scores if there are more than this number. The default is 20 for read mapping. | ||
Revision as of 10:24, 23 June 2014
The data is used in the paper "Count-based differential ....." by Anders et al 2013.
Required tools
The version number indicated below is the one I use. It may be updated when you are ready to download that.
- bowtie (bowtie2-2.1.0-linux-x86_64.zip): Mac, Linux, Windows binaries
- samtools (samtools-0.1.19.tar.bz2): require GCC to compile (build-essential) & zlib1g-dev & libncurses5-dev packages. wikipedia.
- tophat (tophat-2.0.10.Linux_x86_64.tar.gz): Linux and Mac binaries
- sra toolkit (sratoolkit.2.3.4-2-ubuntu64.tar.gz): Mac, Ubuntu & CentOS binaries
- HTseq (HTSeq-0.6.1.tar.gz): require Python 2
- IGV (IGV_2.3.26.zip): require Java and registration to download the program
- FastQC (fastqc_v0.10.1.zip): require Java (jdk is needed, jre is not enough)
- FastX (fastx_toolkit_0.0.13_binaries_Linux_2.6_amd64.tar.bz2): require libgtextutils-0.6
Installation
If binary executables are available, we don't need to do anything special except adding the path containing the binary to the PATH variable.
If only source code is available, make sure the required tool GCC is installed (Under Ubuntu, we can use sudo apt-get install build-essential). Then we can compile and then install the program by using ./configure, make and make install.
In this exercise, all source code or binary programs were extracted to $HOME/binary/ directory (Use mkdir ~/binary to create this new directory) where each program has its own directory.
To use a shell script to install every thing, try the following.
cd; wget -N https://www.dropbox.com/s/ip1jiarxhq0dq91/install.sh; chmod +x install.sh; sudo ./install.sh; rm install.sh
Change PATH variable
Open ~/.basrhc file using any text editor (such as nano or vi) and add the path containing the binary of each program to PATH variable.
tail ~/.bashrc export PATH=$PATH:~/binary/bowtie2-2.1.0/ export PATH=$PATH:~/binary/tophat-2.0.10.Linux_x86_64/ export PATH=$PATH:~/binary/samtools-0.1.19/ export PATH=$PATH:~/binary/samtools-0.1.19/bcftools/ export PATH=$PATH:~/binary/samtools-0.1.19/misc/ export PATH=$PATH:~/binary/sratoolkit.2.3.4-2-ubuntu64/bin export PATH=$PATH:~/binary/HTSeq-0.6.1/build/scripts-2.7 export PATH=$PATH:~/binary/IGV_2.3.26/ export PATH=$PATH:~/binary/FastQC/ export PATH=$PATH:~/binary/fastx/bin
If we want to make these tools available for ALL users (ie system wide environment), we need to put these lines on /etc/profile file.
Data directory
Download <nprot.2013.099-S1.zip> from the paper's web site and extract it to ~/Anders2013 directory.
brb@brbweb4:~$ mkdir Anders2013 brb@brbweb4:~$ cd Anders2013 brb@brbweb4:~/Anders2013$ wget http://www.nature.com/nprot/journal/v8/n9/extref/nprot.2013.099-S1.zip brb@brbweb4:~/Anders2013$ unzip nprot.2013.099-S1.zip brb@brbweb4:~/Anders2013$ ls CG8144_RNAi-1.count counts.csv README.txt Untreated-1.count Untreated-6.count CG8144_RNAi-3.count __MACOSX samples.csv Untreated-3.count CG8144_RNAi-4.count nprot.2013.099-S1.zip SraRunInfo.csv Untreated-4.count
The raw data is from GSE18508 'modENCODE Drosophila RNA Binding Protein RNAi RNA-Seq Studies'.
Subdirectory
During running tophat program, several subdirectories will be generated. Each subdirectory will contain accepted_hits.bam, junctions.bed, insertions.bed, deletions.bed and other files/sub-subdirectories.
Once samtools program was run, .bam (and .bai) files will be created under Anders2013 directory.
Pipeline
The following summary is extracted from http://www.bioconductor.org/help/course-materials/2013/CSAMA2013/tuesday/afternoon/DESeq_protocol.pdf. It is the same as Anders 2013 paper but the paper requires an access permission.
It would be interesting to monitor the disk space usage before, during and after the execution.
Download example data (22 files in SRA format)
Download Supplement file and read <SraRunInfo.csv> in R. We will choose a subset of data based on "LibraryName" column.
sri = read.csv("SraRunInfo.csv", stringsAsFactors=FALSE)
keep = grep("CG8144|Untreated-",sri$LibraryName)
sri = sri[keep,]
fs = basename(sri$download_path)
for(i in 1:nrow(sri))
download.file(sri$download_path[i], fs[i])
names(sri)
[1] "Run" "ReleaseDate" "spots" "bases"
[5] "avgLength" "size_MB" "AssemblyName" "download_path"
[9] "Experiment" "LibraryName" "LibraryStrategy" "LibrarySelection"
[13] "LibrarySource" "LibraryLayout" "InsertSize" "Platform"
[17] "Model" "SRAStudy" "BioProject" "Study_Pubmed_id"
[21] "ProjectID" "Sample" "BioSample" "TaxID"
[25] "ScientificName" "SampleName" "Submission" "Consent"
apply(sri,2, function(x) length(unique(x)))
Run ReleaseDate spots bases
22 22 22 22
avgLength size_MB AssemblyName download_path
5 22 1 22
Experiment LibraryName LibraryStrategy LibrarySelection
7 7 1 1
LibrarySource LibraryLayout InsertSize Platform
1 2 2 1
Model SRAStudy BioProject Study_Pubmed_id
1 1 1 1
ProjectID Sample BioSample TaxID
1 7 1 1
ScientificName SampleName Submission Consent
1 1 1 1
sri[1:2,]
Run ReleaseDate spots bases avgLength size_MB
1 SRR031714 2010-01-15 10:42:00 5327425 394229450 74 396
2 SRR031715 2010-01-21 14:24:18 5248396 388381304 74 390
AssemblyName
1 NA
2 NA
download_path
1 ftp://ftp-private.ncbi.nlm.nih.gov/sra/sra-instant/reads/ByRun/sra/SRR/SRR031/SRR031714/SRR031714.sra
2 ftp://ftp-private.ncbi.nlm.nih.gov/sra/sra-instant/reads/ByRun/sra/SRR/SRR031/SRR031715/SRR031715.sra
Experiment LibraryName LibraryStrategy LibrarySelection LibrarySource
1 SRX014459 Untreated-3 RNA-Seq cDNA TRANSCRIPTOMIC
2 SRX014459 Untreated-3 RNA-Seq cDNA TRANSCRIPTOMIC
LibraryLayout InsertSize Platform Model SRAStudy
1 PAIRED 200 ILLUMINA Illumina Genome Analyzer II SRP001537
2 PAIRED 200 ILLUMINA Illumina Genome Analyzer II SRP001537
BioProject Study_Pubmed_id ProjectID Sample BioSample
1 GEO Series accession: GSE18508 20921232 168994 SRS008447 NA
2 GEO Series accession: GSE18508 20921232 168994 SRS008447 NA
TaxID ScientificName SampleName Submission Consent
1 7227 Drosophila melanogaster Drosophila melanogaster SRA010243 public
2 7227 Drosophila melanogaster Drosophila melanogaster SRA010243 public
Convert SRA to FASTQ (sra -> fastq)
This takes a long time and it is not to run in parallel by default. At the end of execution, there are 30 fastq files because 8 SRA file are pair end.
stopifnot( all(file.exists(fs)) ) # assure FTP download was successful
for(f in fs) {
cmd = paste("fastq-dump --split-3", f)
cat(cmd,"\n")
system(cmd) # invoke command
}
fastq-dump --split-3 SRR031714.sra
Read 5327425 spots for SRR031714.sra
Written 5327425 spots for SRR031714.sra
fastq-dump --split-3 SRR031715.sra
Read 5248396 spots for SRR031715.sra
Written 5248396 spots for SRR031715.sra
...
Note that '--split-3' option will splits mate-pair reads into separate files (First biological reads satisfying dumping conditions are placed in files *_1.fastq and *_2.fastq If only one biological read is present it is placed in *.fastq Biological reads and above are ignored). See this post.
brb@brbweb4:~/Anders2013$ ls -lh *.fastq -rw-r--r-- 1 brb brb 1.3G Feb 10 12:47 SRR031708.fastq -rw-r--r-- 1 brb brb 812M Feb 10 12:50 SRR031709.fastq -rw-r--r-- 1 brb brb 1.2G Feb 10 12:56 SRR031710.fastq -rw-r--r-- 1 brb brb 1.2G Feb 10 13:02 SRR031711.fastq -rw-r--r-- 1 brb brb 1.3G Feb 10 13:08 SRR031712.fastq -rw-r--r-- 1 brb brb 1.3G Feb 10 13:14 SRR031713.fastq -rw-r--r-- 1 brb brb 1.1G Feb 10 11:48 SRR031714_1.fastq -rw-r--r-- 1 brb brb 1.1G Feb 10 11:48 SRR031714_2.fastq -rw-r--r-- 1 brb brb 1.1G Feb 10 11:54 SRR031715_1.fastq -rw-r--r-- 1 brb brb 1.1G Feb 10 11:54 SRR031715_2.fastq -rw-r--r-- 1 brb brb 1.2G Feb 10 12:02 SRR031716_1.fastq -rw-r--r-- 1 brb brb 1.2G Feb 10 12:02 SRR031716_2.fastq -rw-r--r-- 1 brb brb 1.3G Feb 10 12:10 SRR031717_1.fastq -rw-r--r-- 1 brb brb 1.3G Feb 10 12:10 SRR031717_2.fastq -rw-r--r-- 1 brb brb 1.4G Feb 10 13:21 SRR031718.fastq -rw-r--r-- 1 brb brb 927M Feb 10 13:25 SRR031719.fastq -rw-r--r-- 1 brb brb 1.3G Feb 10 13:32 SRR031720.fastq -rw-r--r-- 1 brb brb 1.3G Feb 10 13:39 SRR031721.fastq -rw-r--r-- 1 brb brb 1.2G Feb 10 13:45 SRR031722.fastq -rw-r--r-- 1 brb brb 842M Feb 10 13:49 SRR031723.fastq -rw-r--r-- 1 brb brb 1.2G Feb 10 12:17 SRR031724_1.fastq -rw-r--r-- 1 brb brb 1.2G Feb 10 12:17 SRR031724_2.fastq -rw-r--r-- 1 brb brb 1.3G Feb 10 12:25 SRR031725_1.fastq -rw-r--r-- 1 brb brb 1.3G Feb 10 12:25 SRR031725_2.fastq -rw-r--r-- 1 brb brb 1.3G Feb 10 12:33 SRR031726_1.fastq -rw-r--r-- 1 brb brb 1.3G Feb 10 12:33 SRR031726_2.fastq -rw-r--r-- 1 brb brb 1.2G Feb 10 12:41 SRR031727_1.fastq -rw-r--r-- 1 brb brb 1.2G Feb 10 12:41 SRR031727_2.fastq -rw-r--r-- 1 brb brb 1.3G Feb 10 13:54 SRR031728.fastq -rw-r--r-- 1 brb brb 3.2G Feb 10 14:07 SRR031729.fastq
Access quality control through ShortRead or FastQC
library("ShortRead")
fqQC = qa(dirPath=".", pattern=".fastq$", type="fastq")
report(fqQC, type="html", dest="fastqQAreport")
Use a web browser to inspect the generated HTML file (here, stored in the \fastqQAreport" directory) with the quality-assessment report.
FastX_trimmer is a better choice. The syntax is
fastx_trimmer -f N -l N -i INPUT -o OUTPUT -v -Q33
The last parameter -Q33 is necessary for Illumina encoding quality score. If we omit it, we will get an error fastx_trimmer: Invalid quality score value (char '#' ord 35 quality value -29) on line 4
Download reference genome (for Tophat)
wget ftp://ftp.ensembl.org/pub/release-70/fasta/drosophila_melanogaster/dna/Drosophila_melanogaster.BDGP5.70.dna.toplevel.fa.gz gunzip Drosophila_melanogaster.BDGP5.70.dna.toplevel.fa.gz
Note that bowtie2/tophat2 indices for many commonly used reference genomes can be downloaded directly from http://tophat.cbcb.umd.edu/igenomes.html.
Structure of downloaded reference genome from Tophat
Downloaded genome from tophat takes a long time (20GB for human). Using bowtie2-build program which can takes an enormous time to run.
$ tar xzvf Homo_sapiens_Ensembl_GRCh37.tar.gz
$ tree -L 4 Homo_sapiens
Homo_sapiens
└── Ensembl
└── GRCh37
├── Annotation
│ ├── Archives <==== gtf file
│ ├── Genes -> Archives/archive-current/Genes
│ ├── README.txt -> Archives/archive-current/README.txt
│ ├── SmallRNA -> Archives/archive-current/SmallRNA
│ └── Variation -> Archives/archive-current/Variation
├── GenomeStudio
│ ├── Archives
│ ├── Homo_sapiens -> Archives/archive-2012-03-09-04-49-46/Homo_sapiens
│ └── README.txt
└── Sequence
├── AbundantSequences
├── Bowtie2Index <==== bt2 files
├── BowtieIndex
├── BWAIndex
├── Chromosomes
├── Squashed-Homo_sapiens-Ensembl-GRCh37
└── WholeGenomeFasta <==== fa file
18 directories, 2 files
$ ls -l Homo_sapiens/Ensembl/GRCh37/Annotation/Archives/archive-2013-03-06-14-23-04/Genes
total 639136
-rwxrwxr-x 1 brb brb 302 Mar 18 2013 ChromInfo.txt
-rwxrwxr-x 1 brb brb 603096976 Mar 18 2013 genes.gtf
-rwxrwxr-x 1 brb brb 8236175 Mar 18 2013 refFlat.txt.gz
-rwxrwxr-x 1 brb brb 43130092 Mar 18 2013 refGene.txt
Download GTF (gene transfer format) file (for Tophat)
The compressed file is 77.6 MB size.
wget ftp://ftp.ensembl.org/pub/release-70/gtf/drosophila_melanogaster/Drosophila_melanogaster.BDGP5.70.gtf.gz gunzip Drosophila_melanogaster.BDGP5.70.gtf.gz
CRITICAL: Make sure that the gene annotation uses the same coordinate system as the reference FASTA le. Here, both les use BDGP5 (i. e., release 5 of the assembly provided by the Berkeley Drosophila Genome Project), as is apparent from the le names. To be on the safe side here, we recommend to always download the FASTA reference sequence and the GTF annotation data from the same resource provider.
- Ensembl.org provides both fasta and gtf files together. It's useful when we want to use Galaxy.
- Broad also provides a link to download gtf and fasta file.
Preprocess reference FASTA files into an index (fasta -> bt2)
Alternatively, pre-built indices for many commonly-used genomes are available from http://tophat.cbcb.umd.edu/igenomes.html <Drosophila_melanogaster_Ensembl_BDGP5.tar.gz> contains genome.1.bt2, genome.2.bt2, genome.3.bt2, genome.fa, genome.rev.1.bt2, and genome.rev.2.bt2 under Drosophila_melanogaster/Ensembl/BDGP5/Sequence/Bowtie2Index/ directory. This takes about 4 minutes 37 seconds on my Core-i7 Laptop computer running xubuntu 12.04.
When I try to run bowtie2-build on mouse genome, it is extremely slow. Unfortunately bowtie2-build does not have "-p" option like bowtie2 to speed it up.
bowtie2-build -f Drosophila_melanogaster.BDGP5.70.dna.toplevel.fa Dme1_BDGP5_70 brb@brbweb4:~/Anders2013$ ls -lt *.bt2 -rw-r--r-- 1 brb brb 58770281 Feb 10 12:06 Dme1_BDGP5_70.rev.1.bt2 -rw-r--r-- 1 brb brb 40591960 Feb 10 12:06 Dme1_BDGP5_70.rev.2.bt2 -rw-r--r-- 1 brb brb 58770281 Feb 10 12:04 Dme1_BDGP5_70.1.bt2 -rw-r--r-- 1 brb brb 40591960 Feb 10 12:04 Dme1_BDGP5_70.2.bt2 -rw-r--r-- 1 brb brb 339200 Feb 10 12:01 Dme1_BDGP5_70.3.bt2 -rw-r--r-- 1 brb brb 40591953 Feb 10 12:01 Dme1_BDGP5_70.4.bt2
Create a metadata table called "samples"
sri$LibraryName = gsub("S2_DRSC_","",sri$LibraryName) # trim label
samples = unique(sri[,c("LibraryName","LibraryLayout")])
for(i in seq_len(nrow(samples))) {
rw = (sri$LibraryName==samples$LibraryName[i])
if(samples$LibraryLayout[i]=="PAIRED") {
samples$fastq1[i] = paste0(sri$Run[rw],"_1.fastq",collapse=",")
samples$fastq2[i] = paste0(sri$Run[rw],"_2.fastq",collapse=",")
} else {
samples$fastq1[i] = paste0(sri$Run[rw],".fastq",collapse=",")
samples$fastq2[i] = ""
}
}
samples$condition = "CTL"
samples$condition[grep("RNAi",samples$LibraryName)] = "KD"
samples$shortname = paste( substr(samples$condition,1,2),
substr(samples$LibraryLayout,1,2), seq_len(nrow(samples)), sep=".")
samples
LibraryName LibraryLayout
1 Untreated-3 PAIRED
3 Untreated-4 PAIRED
5 CG8144_RNAi-3 PAIRED
7 CG8144_RNAi-4 PAIRED
144 Untreated-1 SINGLE
150 CG8144_RNAi-1 SINGLE
156 Untreated-6 SINGLE
fastq1
1 SRR031714_1.fastq,SRR031715_1.fastq
3 SRR031716_1.fastq,SRR031717_1.fastq
5 SRR031724_1.fastq,SRR031725_1.fastq
7 SRR031726_1.fastq,SRR031727_1.fastq
144 SRR031708.fastq,SRR031709.fastq,SRR031710.fastq,SRR031711.fastq,SRR031712.fastq,SRR031713.fastq
150 SRR031718.fastq,SRR031719.fastq,SRR031720.fastq,SRR031721.fastq,SRR031722.fastq,SRR031723.fastq
156 SRR031728.fastq,SRR031729.fastq
fastq2 condition shortname
1 SRR031714_2.fastq,SRR031715_2.fastq CTL CT.PA.1
3 SRR031716_2.fastq,SRR031717_2.fastq CTL CT.PA.2
5 SRR031724_2.fastq,SRR031725_2.fastq KD KD.PA.3
7 SRR031726_2.fastq,SRR031727_2.fastq KD KD.PA.4
144 CTL CT.SI.5
150 KD KD.SI.6
156 CTL CT.SI.7
Align the reads to the reference genome using tophat2 (fastq -> bam, bed)
Note:
- '-p' specifies the number of threads to use, -o specifies the output directory. The first name (bowind) is the name of the index (built in advance).
- It seems even the program cannot find the reference genome, it still runs.
- The run time is about 3 to 4 hours per sample when 7 jobs were running at the same time (5 threads for each sample).
- If only one sample was run it took 45 minutes as the paper said (5 threads). If I specify 1 thread, the running time is 1 hour 44 minutes.
- If I only run 2 jobs using batch, it takes 1 hour and 1 hour 2 mintues to finish these two jobs.
- The input is fastq files. The output is a directory contains 7 files (including *.bam) and 1 subdirectory of logs.
gf = "Drosophila_melanogaster.BDGP5.70.gtf"
bowind = "Dme1_BDGP5_70"
cmd = with(samples, paste("tophat2 -G", gf, "-p 5 -o", LibraryName,
bowind, fastq1, fastq2))
cmd # Actually we want to use system() to run each line of cmd.
[1] "tophat2 -G Drosophila_melanogaster.BDGP5.70.gtf -p 5 -o Untreated-3 Dme1_BDGP5_70 SRR031714_1.fastq,SRR031715_1.fastq SRR031714_2.fastq,SRR031715_2.fastq"
[2] "tophat2 -G Drosophila_melanogaster.BDGP5.70.gtf -p 5 -o Untreated-4 Dme1_BDGP5_70 SRR031716_1.fastq,SRR031717_1.fastq SRR031716_2.fastq,SRR031717_2.fastq"
[3] "tophat2 -G Drosophila_melanogaster.BDGP5.70.gtf -p 5 -o CG8144_RNAi-3 Dme1_BDGP5_70 SRR031724_1.fastq,SRR031725_1.fastq SRR031724_2.fastq,SRR031725_2.fastq"
[4] "tophat2 -G Drosophila_melanogaster.BDGP5.70.gtf -p 5 -o CG8144_RNAi-4 Dme1_BDGP5_70 SRR031726_1.fastq,SRR031727_1.fastq SRR031726_2.fastq,SRR031727_2.fastq"
[5] "tophat2 -G Drosophila_melanogaster.BDGP5.70.gtf -p 5 -o Untreated-1 Dme1_BDGP5_70 SRR031708.fastq,SRR031709.fastq,SRR031710.fastq,SRR031711.fastq,SRR031712.fastq,SRR031713.fastq "
[6] "tophat2 -G Drosophila_melanogaster.BDGP5.70.gtf -p 5 -o CG8144_RNAi-1 Dme1_BDGP5_70 SRR031718.fastq,SRR031719.fastq,SRR031720.fastq,SRR031721.fastq,SRR031722.fastq,SRR031723.fastq "
[7] "tophat2 -G Drosophila_melanogaster.BDGP5.70.gtf -p 5 -o Untreated-6 Dme1_BDGP5_70 SRR031728.fastq,SRR031729.fastq "
Tophat options
- --coverage-search/--no-coverage-search: According to this, it is a step to define possible junctions between exons. If you only want the expression profile of the annotated genes, you can skip this step for speed. The identification of new regions with the coverage is relevant only if you want to detect new splice sites in alternate transcripts or even new genes.
- -r/--mate-inner-dist <int>: This is the expected (mean) inner distance between mate pairs. For, example, for paired end runs with fragments selected at 300bp, where each end is 50bp, you should set -r to be 200. The default is 50bp.
- -g/--max-multihits <int>: Instructs TopHat to allow up to this many alignments to the reference for a given read, and choose the alignments based on their alignment scores if there are more than this number. The default is 20 for read mapping.
- --library-type: The default is unstranded (fr-unstranded). If either fr-firststrand or fr-secondstrand is specified, every read alignment will have an XS attribute tag as explained below. Consider supplying library type options below to select the correct RNA-seq protocol.
Output from console
[2014-04-18 16:42:12] Beginning TopHat run (v2.0.10) ----------------------------------------------- [2014-04-18 16:42:12] Checking for Bowtie Bowtie version: 2.1.0.0 [2014-04-18 16:42:12] Checking for Samtools Samtools version: 0.1.19.0 [2014-04-18 16:42:12] Checking for Bowtie index files (genome).. [2014-04-18 16:42:12] Checking for reference FASTA file [2014-04-18 16:42:12] Generating SAM header for genome [2014-04-18 16:42:58] Reading known junctions from GTF file [2014-04-18 16:43:17] Preparing reads left reads: min. length=49, max. length=49, 26721504 kept reads (2587 discarded) [2014-04-18 16:47:40] Building transcriptome data files E2_Rep2/tmp/genes [2014-04-18 16:48:22] Building Bowtie index from genes.fa [2014-04-18 17:12:33] Mapping left_kept_reads to transcriptome genes with Bowtie2 [2014-04-18 17:44:09] Resuming TopHat pipeline with unmapped reads [2014-04-18 17:44:09] Mapping left_kept_reads.m2g_um to genome genome with Bowtie2 [2014-04-18 17:49:50] Mapping left_kept_reads.m2g_um_seg1 to genome genome with Bowtie2 (1/2) [2014-04-18 17:51:07] Mapping left_kept_reads.m2g_um_seg2 to genome genome with Bowtie2 (2/2) [2014-04-18 17:51:54] Searching for junctions via segment mapping Coverage-search algorithm is turned on, making this step very slow Please try running TopHat again with the option (--no-coverage-search) if this step takes too much time or memory. [2014-04-18 18:08:13] Retrieving sequences for splices [2014-04-18 18:10:10] Indexing splices [2014-04-18 18:19:15] Mapping left_kept_reads.m2g_um_seg1 to genome segment_juncs with Bowtie2 (1/2) [2014-04-18 18:21:26] Mapping left_kept_reads.m2g_um_seg2 to genome segment_juncs with Bowtie2 (2/2) [2014-04-18 18:23:38] Joining segment hits [2014-04-18 18:25:52] Reporting output tracks ----------------------------------------------- [2014-04-18 18:38:05] A summary of the alignment counts can be found in E2_Rep2/align_summary.txt [2014-04-18 18:38:05] Run complete: 01:55:52 elapsed
Organize, sort and index the BAM files and create SAM files (bam -> sam)
Note
- Potentially we can run the commands in parallel. This step needs lots of disk I/O so it may be better not to run the command in parallel.
- Four steps:
- first step - Input: original aligned bam file. Output: <Untreated-3_sn.bam> (1.5 GB). sort alignment file by read name (-n option).
- Second step - Input: sorted bam file from 1st step. Output: <Untreated-3_sn.sam> (11 GB). BAM<->SAM conversion. The SAM file generated from these 2 steps will be used in HTSEQ-count.
- Third step - Input: original aligned bam file. Output: <Untreated-3_s.bam> (1.2 GB). sort alignment file by chromosomal coordinates.
- Fourth step - Input: sorted bam file from 3rd step. Output: <Untreated-3_s.bam.bai> (375 KB). index alignment. The SAM and BAI files generated from these 2 steps together with GTF file will be used in IGV.
- Examples how to use. See here
for(i in seq_len(nrow(samples))) {
lib = samples$LibraryName[i]
ob = file.path(lib, "accepted_hits.bam")
# sort by name, convert to SAM for htseq-count
cat(paste0("samtools sort -n ",ob," ",lib,"_sn"),"\n")
cat(paste0("samtools view -o ",lib,"_sn.sam ",lib,"_sn.bam"),"\n")
# sort by position and index for IGV
cat(paste0("samtools sort ",ob," ",lib,"_s"),"\n")
cat(paste0("samtools index ",lib,"_s.bam"),"\n\n")
}
samtools sort -n Untreated-3/accepted_hits.bam Untreated-3_sn
samtools view -o Untreated-3_sn.sam Untreated-3_sn.bam
samtools sort Untreated-3/accepted_hits.bam Untreated-3_s
samtools index Untreated-3_s.bam
samtools sort -n Untreated-4/accepted_hits.bam Untreated-4_sn
samtools view -o Untreated-4_sn.sam Untreated-4_sn.bam
samtools sort Untreated-4/accepted_hits.bam Untreated-4_s
samtools index Untreated-4_s.bam
samtools sort -n CG8144_RNAi-3/accepted_hits.bam CG8144_RNAi-3_sn
samtools view -o CG8144_RNAi-3_sn.sam CG8144_RNAi-3_sn.bam
samtools sort CG8144_RNAi-3/accepted_hits.bam CG8144_RNAi-3_s
samtools index CG8144_RNAi-3_s.bam
samtools sort -n CG8144_RNAi-4/accepted_hits.bam CG8144_RNAi-4_sn
samtools view -o CG8144_RNAi-4_sn.sam CG8144_RNAi-4_sn.bam
samtools sort CG8144_RNAi-4/accepted_hits.bam CG8144_RNAi-4_s
samtools index CG8144_RNAi-4_s.bam
samtools sort -n Untreated-1/accepted_hits.bam Untreated-1_sn
samtools view -o Untreated-1_sn.sam Untreated-1_sn.bam
samtools sort Untreated-1/accepted_hits.bam Untreated-1_s
samtools index Untreated-1_s.bam
samtools sort -n CG8144_RNAi-1/accepted_hits.bam CG8144_RNAi-1_sn
samtools view -o CG8144_RNAi-1_sn.sam CG8144_RNAi-1_sn.bam
samtools sort CG8144_RNAi-1/accepted_hits.bam CG8144_RNAi-1_s
samtools index CG8144_RNAi-1_s.bam
samtools sort -n Untreated-6/accepted_hits.bam Untreated-6_sn
samtools view -o Untreated-6_sn.sam Untreated-6_sn.bam
samtools sort Untreated-6/accepted_hits.bam Untreated-6_s
samtools index Untreated-6_s.bam
Output from console
[bam_sort_core] merging from 15 files...
Inspect alignments with IGV
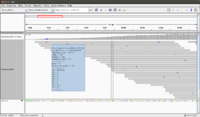
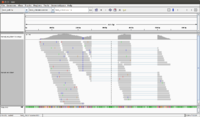
The following screenshot was taken from another (small) dataset. There are 100 sequences in the fq file. The reference genome has only 13 sequences. See also this post.
To use IGV, we have to run samtools twice (one is to get sorted bam file, and the other is to get bai file). Both bam and bai files are binary.
- Genome -> Load genome from file -> test_ref.fa
- File -> load from file -> read_1.fq
- Each seq from fa file will be connected together and shown on the bottom of IGV.
- Mouse over a sequence will show "Read name" (we can go back to fastq file to double check the sequence), "Location" (the location from mouse over), "Alignment start" (the start position).
- If there is a mismatch from a bp, it will be shown in a different color (T-red, C-blue, A-green, G-brown).
- The histogram-like plot is called Coverage.
- One sequence may be splitted after alignment.
- In this example, we can count there are 72 sequences. This matches with the file in <align_summary.txt> where it shows the number of reads in input and number of mapped reads with percentage of mapped.
The code is given below
tophat -r 20 test_ref reads_1.fq reads_2.fq samtools sort tophat_out/accepted_hits.bam tophat_out_s samtools index tophat_out_s.bam
The "-r" option is tophat is related to mate-inner-dist. See the discussion here.
Count reads using htseq-count (sam -> count)
samples$countf = paste(samples$LibraryName, "count", sep=".")
gf = "Drosophila_melanogaster.BDGP5.70.gtf"
cmd = paste0("htseq-count -s no -a 10 ",
samples$LibraryName, "_sn.sam ",
gf," > ", samples$countf)
cmd
[1] "htseq-count -s no -a 10 Untreated-3_sn.sam Drosophila_melanogaster.BDGP5.70.gtf > Untreated-3.count"
[2] "htseq-count -s no -a 10 Untreated-4_sn.sam Drosophila_melanogaster.BDGP5.70.gtf > Untreated-4.count"
[3] "htseq-count -s no -a 10 CG8144_RNAi-3_sn.sam Drosophila_melanogaster.BDGP5.70.gtf > CG8144_RNAi-3.count"
[4] "htseq-count -s no -a 10 CG8144_RNAi-4_sn.sam Drosophila_melanogaster.BDGP5.70.gtf > CG8144_RNAi-4.count"
[5] "htseq-count -s no -a 10 Untreated-1_sn.sam Drosophila_melanogaster.BDGP5.70.gtf > Untreated-1.count"
[6] "htseq-count -s no -a 10 CG8144_RNAi-1_sn.sam Drosophila_melanogaster.BDGP5.70.gtf > CG8144_RNAi-1.count"
[7] "htseq-count -s no -a 10 Untreated-6_sn.sam Drosophila_melanogaster.BDGP5.70.gtf > Untreated-6.count"
Output from console
Lots of output
.... 37200000 SAM alignment records processed. 37300000 SAM alignment records processed. 37399340 SAM alignments processed.
Note: Another tool is CuffLinks. See Sean Davis's tutorial.
DESeq for DE analysis
> samplesDESeq = with(samples, data.frame(
shortname = I(shortname),
countf = I(countf),
condition = condition,
LibraryLayout = LibraryLayout))
> samplesDESeq
shortname countf condition LibraryLayout
1 CT.PA.1 Untreated-3.count CTL PAIRED
2 CT.PA.2 Untreated-4.count CTL PAIRED
3 KD.PA.3 CG8144_RNAi-3.count KD PAIRED
4 KD.PA.4 CG8144_RNAi-4.count KD PAIRED
5 CT.SI.5 Untreated-1.count CTL SINGLE
6 KD.SI.6 CG8144_RNAi-1.count KD SINGLE
7 CT.SI.7 Untreated-6.count CTL SINGLE
> library("DESeq")
cds <- newCountDataSetFromHTSeqCount(samplesDESeq) # quick, 15682 features, 7 samples
cds <- estimateSizeFactors(cds) # quick
sizeFactors(cds)
# CT.PA.1 CT.PA.2 KD.PA.3 KD.PA.4 CT.SI.5 KD.SI.6 CT.SI.7
# 0.6991612 0.8104921 0.8217403 0.8941097 1.6431467 1.3720872 1.1041186
cds <- estimateDispersions(cds) # quick
res <- nbinomTest(cds,"CTL","KD") # 44 seconds
sum(res$pval <= .05, na.rm=T) # [1] 1574
sum(res$padj <= .05, na.rm=T) # [1] 730
options(width=100)
res[1:3,]
# id baseMean baseMeanA baseMeanB foldChange log2FoldChange pval padj
# 1 FBgn0000003 0.000000 0.000000 0.000000 NaN NaN NA NA
# 2 FBgn0000008 88.526695 88.486694 88.580029 1.001055 0.001520941 0.9298627 1
# 3 FBgn0000014 3.054418 2.327533 4.023599 1.728697 0.789684778 0.6839858 1
File Format
http://genome.ucsc.edu/FAQ/FAQformat.html which includes a lot of common formats like BAM, BED, bedGraph, bigBed, bigWig, VCF, WIG, et al.
Other RNA-Seq data
GEO
Go to http://www.ncbi.nlm.nih.gov/sites/entrez and select 'GEO Datasets' from the drop down menu, and use 'RNA-Seq' as your search term.
GSE28666
as used in DAFS paper. The timing is
- bowtie2-build 115 minutes
- fastq-dump 25 minutes per fastq (3.3 - 5.9GB fastq files)
- tophat2 90 minutes per fastq
- samtools 5 minutes
- htseq-count 13 minutes
GSE51403
RNA-seq differential expression studies: more sequence, or more replication?
GSE37544
ENCODE
http://www.ncbi.nlm.nih.gov/geo/info/ENCODE.html has a list of data with GSE number.
1000 genomes
Human BodyMap 2.0 data from Illumina
- http://www.ensembl.info/blog/2011/05/24/human-bodymap-2-0-data-from-illumina/
- https://usegalaxy.org/u/jeremy/p/galaxy-rna-seq-analysis-exercise. Galaxy needs to install hg19 genome before we can use tophat or bowtie. See Galaxy/Wiki/Admin/Data Preparation page. Genome can be downloaded from tophat webpage. It is 20GB for hg19! We can also download just chromosome 19 fasta file chr19.fa.gz from UCSC genomes FTP server and it is only 19MB. We can then create reference index files (.bt2) by using bowtie2-build command. If we use Galaxy, we don't need to create reference index files.
Other RNA-Seq tools
http://en.wikipedia.org/wiki/List_of_RNA-Seq_bioinformatics_tools
Galaxy
The first 3 lines will install galaxy on local computer. The last command will start Galaxy. PS. It will take a while to start the first time.
hg clone https://bitbucket.org/galaxy/galaxy-dist/ cd galaxy-dist hg update stable sh run.sh
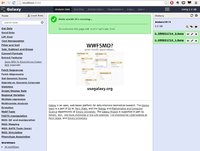
A screenshot of the available tools is given below. Strangely, I did not find "NGS: RNA Analysis" in the NGS: ToolBox on left hand side.
We cannot upload > 2GB fastq data. For such large dataset (we have one 3.3 GB fastq in this case), Galaxy asks to use http or ftp to load the data. I install Apache in Ubuntu and put this fastq file there for upload.
Galaxy does not include bowtie, tophat, .... by default. See this post and Galaxy wiki about the installation. Another option is to modify line 614 of 'universe_wsgl.ini' file and enable ourselves as adminstrator. Also modify line 146 to add a path for installing tools through Galaxy Admin page.
If you submit a lot of jobs (eg each tophat2 generates 5 jobs) some of jobs cannot be run immediately. In my experience, there is a 25 jobs cap. Running jobs has a yellow background color. Waiting for run jobs has a gray background color.
Some helpful information
- https://wiki.galaxyproject.org/CitingGalaxy Citation
- https://wiki.galaxyproject.org/Learn Search rna-seq or NGS as keywords.
- https://biostar.usegalaxy.org/
- Galaxy & DESeq2
Cufflink -> Cuffcompare -> Cuffdiff
- http://cufflinks.cbcb.umd.edu/tutorial.html
- UCSF wiki.
- Michigan State University
- http://seqanswers.com/forums/showthread.php?t=9753
- https://docs.uabgrid.uab.edu/wiki/UAB_Galaxy_RNA_Seq_Step_by_Step_Tutorial#Cufflinks_inputs
- https://sites.google.com/site/princetonhtseq/tutorials/rna-seq
- http://www.genomecenter.ucdavis.edu/resources/instructional-videos